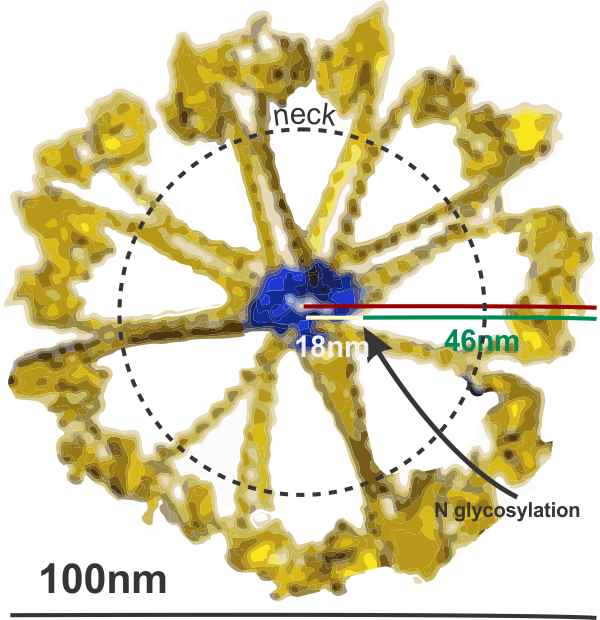
I have found that most of the diagrams of surfactant protein D are pretty far from reality. I found this one however that attempts to be more specific and utilizes at least some of the molecular modeling of that protein available. The RCSB PDB really doesn’t model SP-D in any complete way and while researchers have added many many variations of the carbohydrate recognition domain and the coiled coil neck region, nothing else much comes up as to the rest of the molecule. So here is an attempt (which I have vectorized to eliminate the extraneous stuff in the diagram at least it utilizes the structures known for the trimer — from the neck up (haha). Also, it is totally confusing whether the arms are really trimers, or not, because the shape of the CRDs is relatively large compared to the whole molecule and is not tightly intertwined at the neck region (which is the way the “real” molecule is. So Some of these arms are so spread out that there is no intertwining at the neck region whatsoever. So this is not a good representation of the molecular structure, even though it wold suggest that it is a molecular model. So therein lies a dilemma. 1) to make a diagram so wildly wrong that no one thinks you are making a molecularly accurate representation, or do your best, and make mistakes. One thing for certain…mistakes get perpetuated. 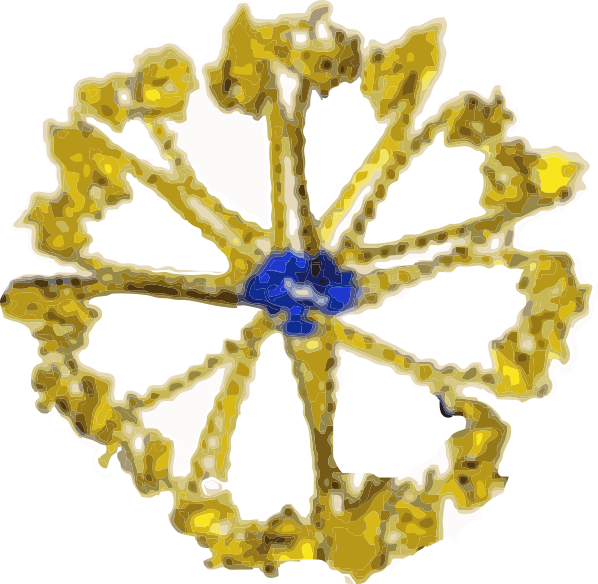
What it does show is the reasonably good approximation of the CRD, and also the curved nature of the SP-D trimeric arms which very few diagrams show but the SP-D trimers and dodecamer molecules seen by TEM show very clearly. What this diagram doesn’t do very well is look at the N terminal portion of SP-D, or show the possibility that there is a connection of the trimers at a “distance” from dead-enter, and it doesn’t indicate where the N terminal glycosylation site might be relative to the whole. Measurements of the latter are taken from a publication by Crouch. Also, it seems that in this particular diagam the collagen-like area here is just a kind of an extended “neck”, with the wrong number of coils, and with no kink, no bend, no separate indication as to where the neck might connect to the collagen-like region. What it also does do, however, is make this particular fuzzyball in a configuration with even numbers of dodecamers (in this case 4, that is 16 arms) which would go along with the TEM images which suggest that the fuzzyballs are an aggregate of dodecamers. What it does not do is look at the possibility of open and closed, or segmented organization of the dodecamers. That would have shown up with more obtuse angles between the four dodecamers and more acute angles within each of the four dodecamer units.
Image found in the following review by Elena N. Atochina-Vasserman
I am suggesting the following structures might have been noted… just by what has been published by Crouch and other researchers on SP-D.