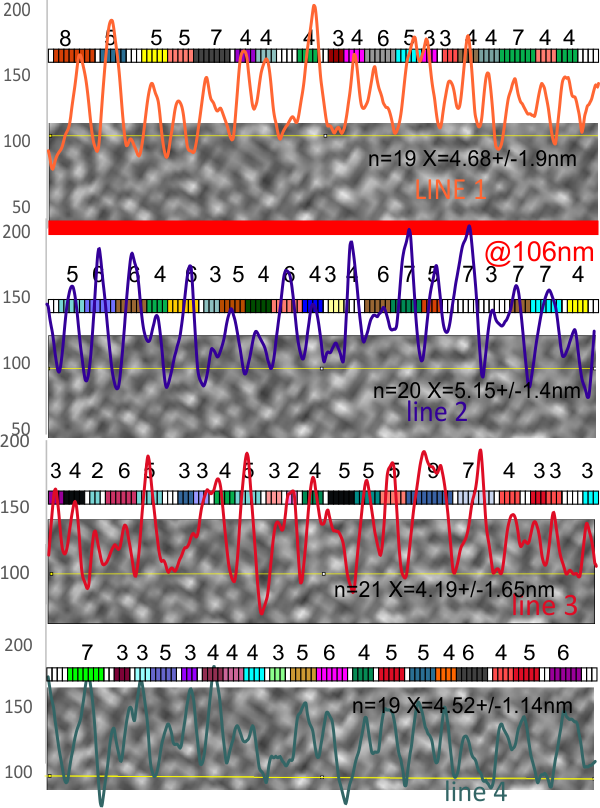
Confirming the background grain size in a particular image (so I can do LUT plots on a few SP-D dodecamers and type 1 collagen molecules for comparisoin with AFM images) is pretty uniform… varying about 1.5 nm (n=79 x=4.63+/-1.45 nm. This means that it might be possible to establish areas of greater luminance (brightness, lightness) on a 1-255 gray scale that correspond with different AA sequences in either (both) these molecules.
Individual lines and their nm thick peak-valleys, and a mean of all 4 lines together. Same plots as last week, same resource, just with the measurements of peak width (from lowest point in valley one side to the other.
and the two LUT plots made with rectangles, and two more plots made with diagonals. It seems that the rectangle might be the least good way to measure the nm of the shadowed background. But all in all, it seems that a line, anyway you draw it will work just fine.
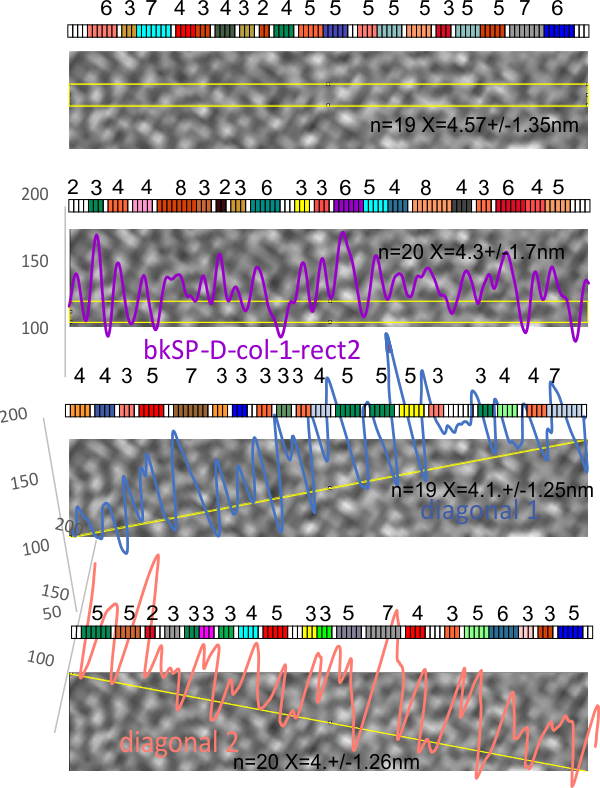 The next step is to see the width and variation in the type 1 collagen collagen and SP-D from the same micrograph.
The next step is to see the width and variation in the type 1 collagen collagen and SP-D from the same micrograph.