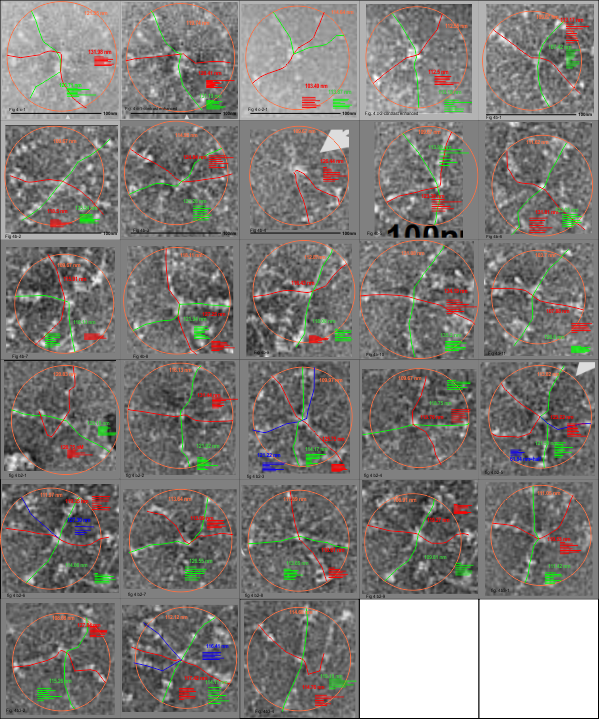
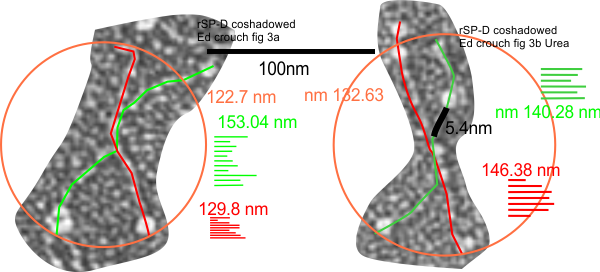
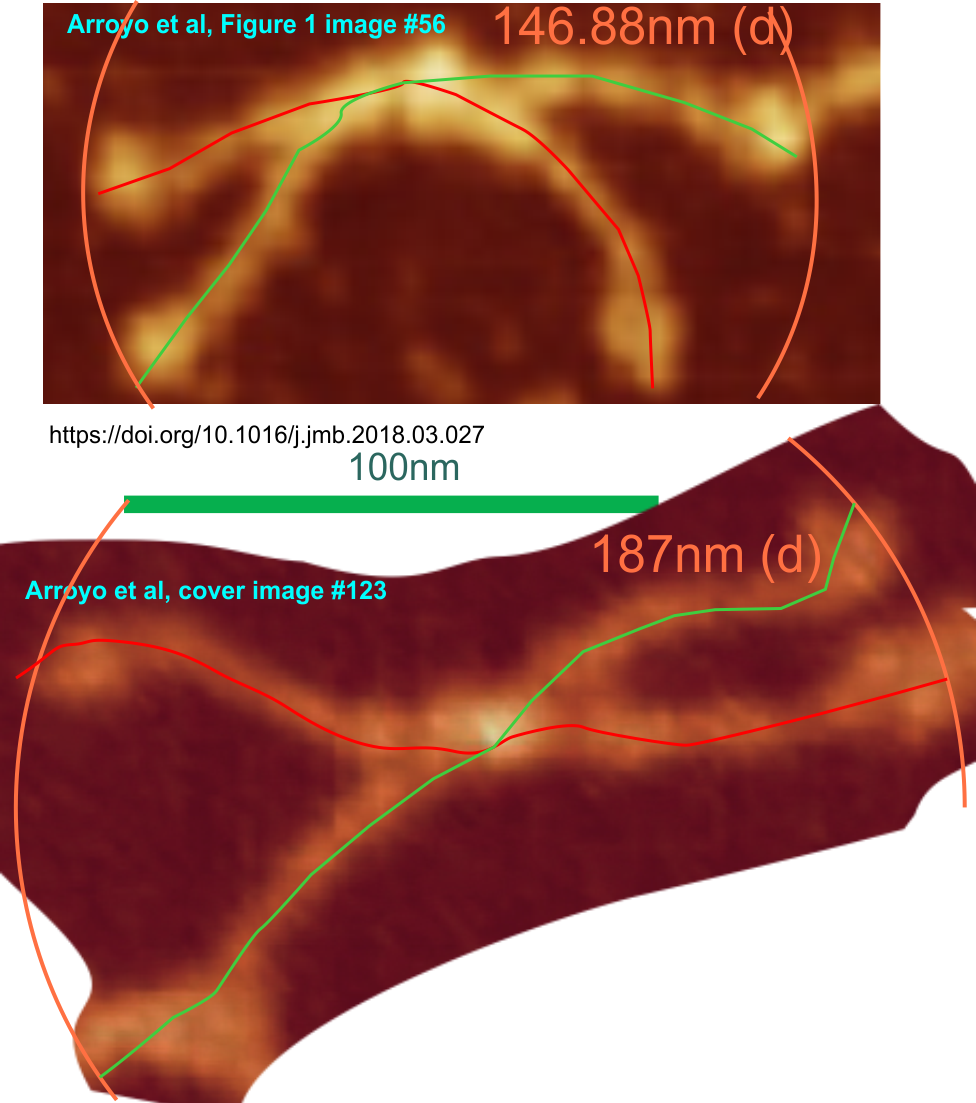
I have measured the size of the SP-D images in the article by Perino et al, and their assessment of whether SP-D assists in controlling infection with vaccinia virus. The purpose was to assess whether there was a difference in the size of the molecules from one figure to the next, (which there ws not : Fig 4c-molecules 1-2 =118.71 nm +/ 7.8 nm; Figure 4b 1-11 =118.73 nm +/ 8.76 nm; Figure 4b2 molecules 1-9; =117.27 nm +/ 5.94 nm; Figure 4b3 molecules 1-4; 115.52 nm +/ 4.74 nm. They provided bar markers for each of the images and this was used to determine the arm lengths and total diameter. (SEE MY figure below).
This is the first negative stained group of SP-D dodecamers and lower number multimers that I have measured using the line-node method and the diameter method. While the text appears to suggest that this is recombinant human SP-D (just like Arroyo et al says their dodecamers are recombinant human SP-D) the difference in size between the AFM (latter paper) and negative staining (Perrino et al is huge) i.e. 30 nm approximately.
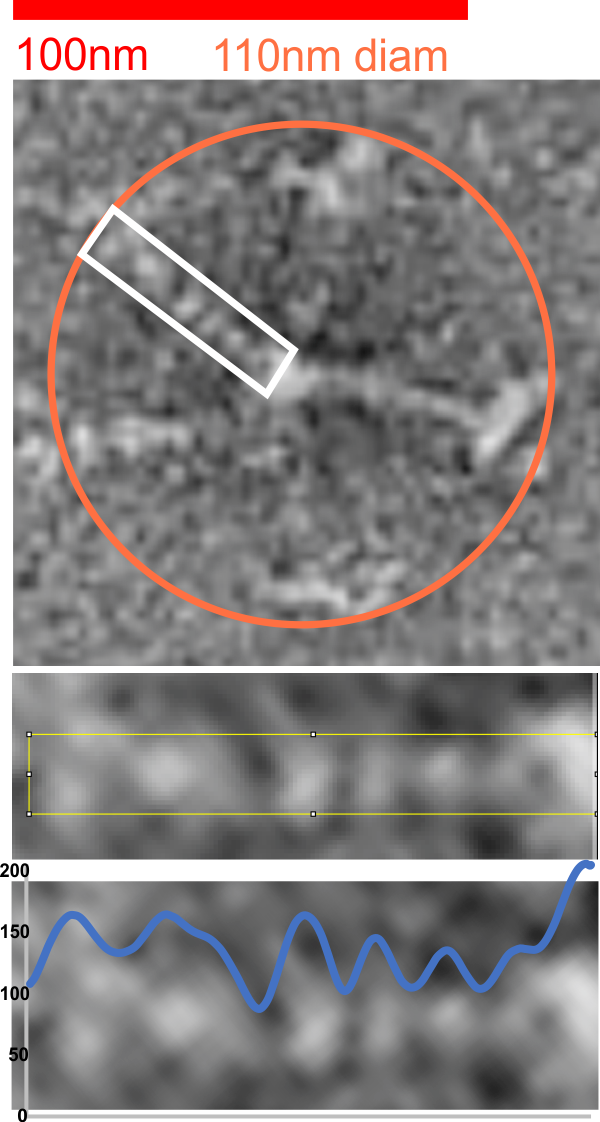
Figure below shows exactly what molecules looked like, and exactly where i anticipated the molecules to begin and end.