Going through archived images of the livers of mice which received infusions of iodo containing perfluorochemicals, I ran onto a hand-written comment which made me laugh. It was “funky print given under pressure to LC Clark (Jr) that perfluorodecyl iodide increases tremendously the number and size of microbodies (aka peroxisomes) in the mouse liver” (words in italics added today for clarity). Apparently no one wanted to hear that news in 1973 or 4… perhaps because it was a “damning” mark on the possible use of perfluorochemical formulations in general, and specifically for artificial – blood substitutes.
Wikipedia summary is here: “They are involved in catabolism of very long chain fatty acids, branched chain fatty acids, D-amino acids, and polyamines, reduction of reactive oxygen species – specifically hydrogen peroxide; – and biosynthesis of plasmalogens, such as ether phospholipids critical for the normal function of mammalian brains and lungs. They also contain approximately 10% of the total activity of two enzymes in the pentose phosphate pathway, which is important for energy metabolism.”
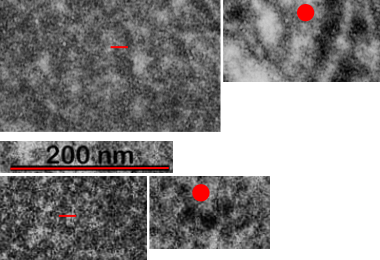
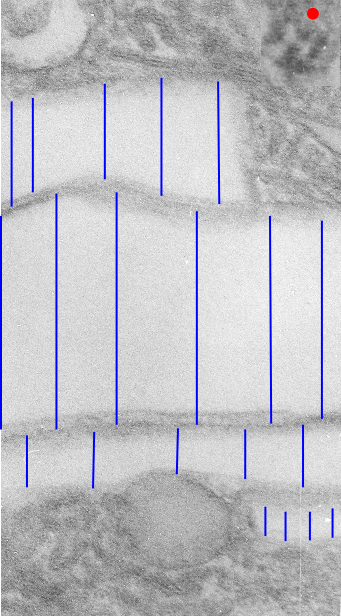
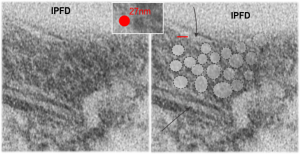
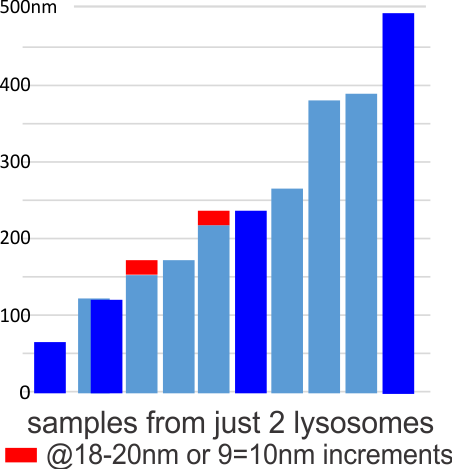
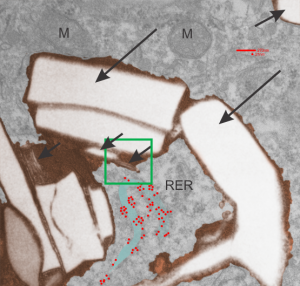
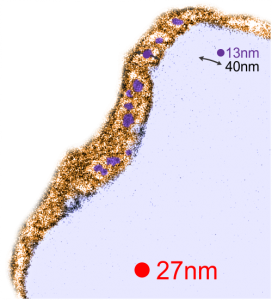
Below are three images, one of liver which shows the very large peroxisomes, in this particular micrograph (not all shown here) they were a volume density of about 30% of the cytoplasm which puts IPFD up there with the best of the proliferators; mitochondria for comparison were about 33% of the cytoplasm. Consensus has peroxisomes at more like 2% of the volume density of the hepatocyte under unstimulated conditions.
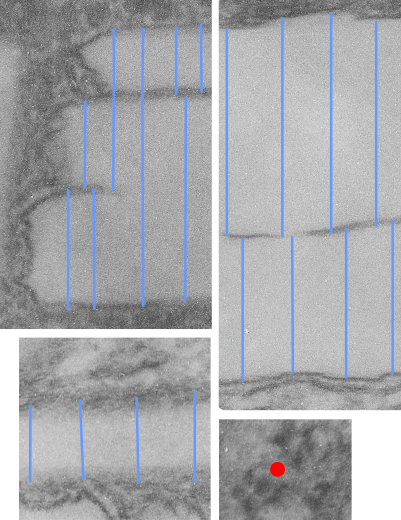
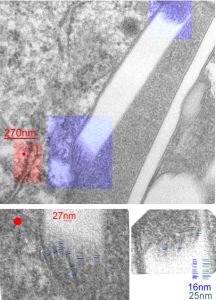
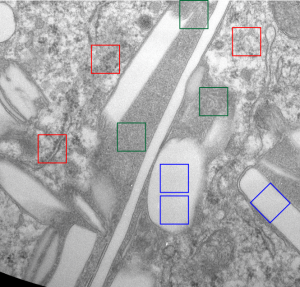
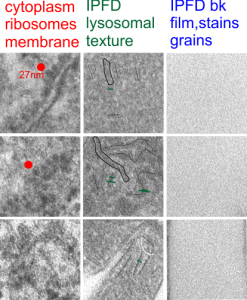
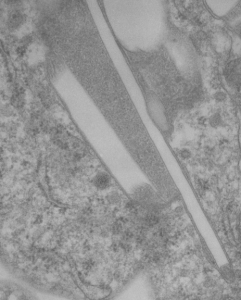
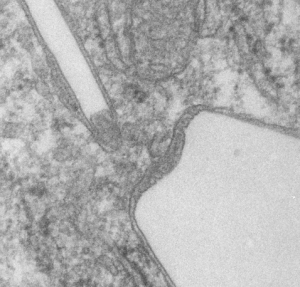
The story will be complicated as there are more than 30 enzymes which are in peroxisomes: and the patterning that is seen in the enzymes within the phagolysosomes containing IPFD looks to be similarly wound as the enzymes within peroxisomes in the hepatocytes (top two image groups). From the internet the middle three images (used without permission as a screen print of the insanely egotistical Dr. Jastro TEM collection who most certainly used public funds for this research and collection of micrographs and yet thinks he has ownership of these images…. not civil at all in my opinion and shame on him) is an hepatocyte peroxisome (which i measured against a nearby ribosome and also the micron bar marker, for size) shows a coarse texture, somewhat coiled. The latter and my peroxisome images (top), added from a portion of an hepatocyte from the liver which received IPFD and measured against a nearby ribosome, show similar “oligomerization” or “substructural” protein patterning.
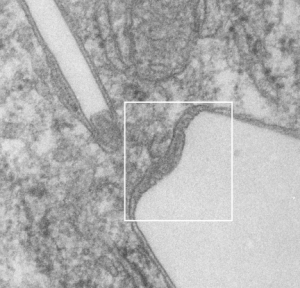
Some protein(s) is creating this texture. It would be interesting to know if the protein in these microbodies (aka peroxisomes) is the same as that which is found in abundance in the phagolysosomes containing IPFD. Bottom micrograph is a portion of hepatocyte cytoplasm from IPFD infused mouse more than 200 days post infusion from which my (top two) images are derived.