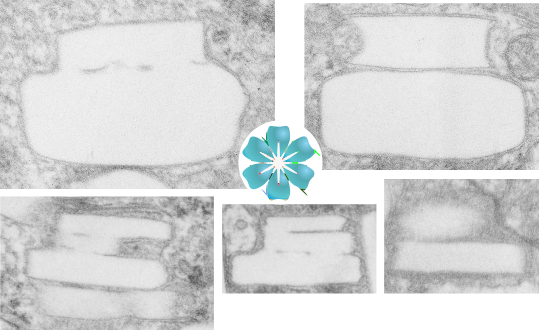
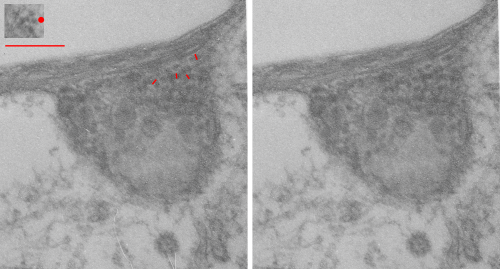
I sometimes get the uneasy feeling that if i see a pattern in an electron micrograph that someone is going to say, ah thats just fixation artifact… and of course they are right. All testing is “fixation artifact” but its the purview of each observer to determine what is background noise and what is real structure.
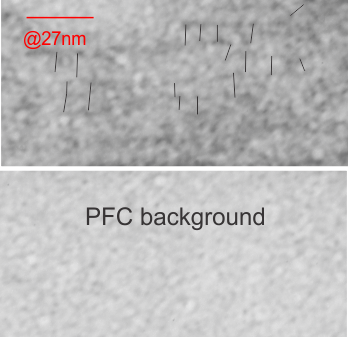
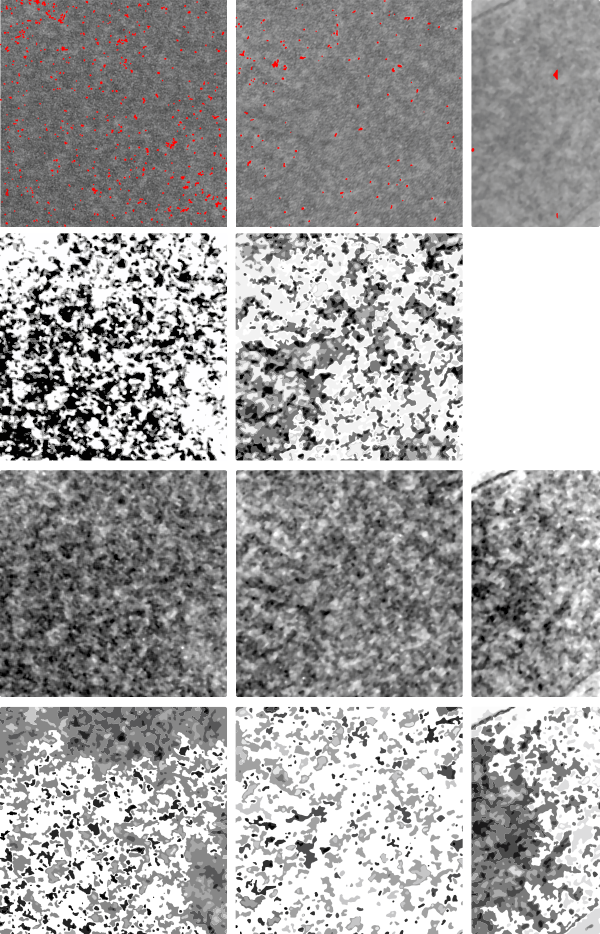
In the case of looking at perfluorochemicals left behind in tissues, consensus says that there is “nothing” in that empty footprint, just the embedding medium (in this case EPON 812). So there is a texture to the embedding medium, no question, but nothing I can perceive, and so easily compared to the texture of surrounding tissues, so that when I see a pattern in the tissue I am pretty convinced that it is the presence of the proteins . (They are influenced by the fixative and type and length of dehydration and other stuff not excluding the warping and shifting under the electron microscope. All those factors play a role). It is one of the reasons that an internal control for size (and i am suggesting background texture) in electron microscopy is really important. I CAN list a short list of variables to be aware of (my knowledge is just observational) which begin even before biopsy, including the time of the last meal, 1) circadian cycle, 2) health of the tissue (in vivo or in vitro –doesn’t matter) 3) mode of analgesia or anesthesia at biopsy (paralytics – solutions with ions (as in perfusion) 4) time till tissue gets into fixative, 5) size of the tissue block, 6) type of fixative, 7) length of time in fixative, 8) length of time and chemicals used for dehydration, 9)type of plastic, 10) section thickness, 11) staining compounds, 12) TEM specs, KV, etc…. so I am willing to accept that not all parameters are under one’s control.
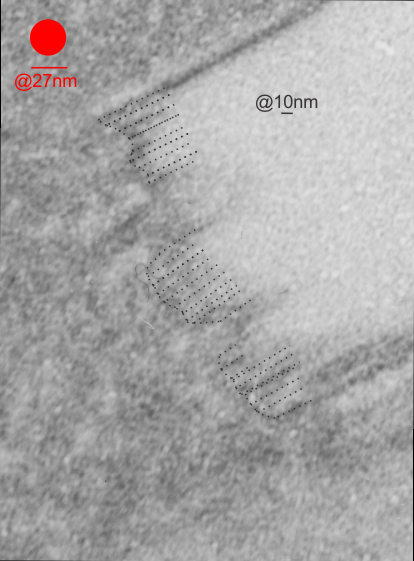
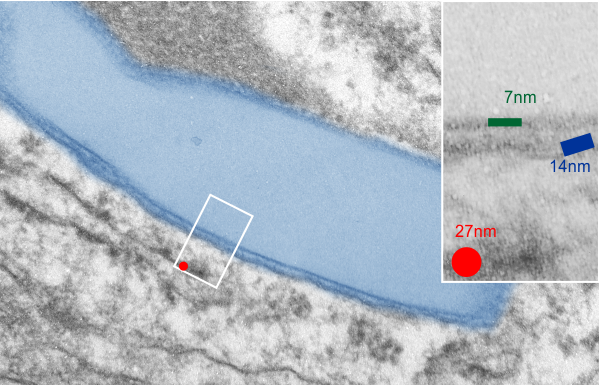
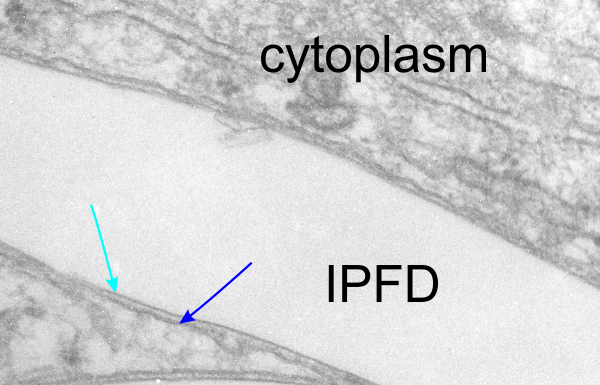
I will however say that the pattern below (in a portion of a macrophage in the liver of a mouse given 10%IPFD in 5%F68, 267 days post infusion) shows a linearity I would not be willing to ignore. low mag micrograph top, for orientation, next below is micrograph of the red box area which shows the periodicity. That is just one of the linear arrangements that are visible in these tissues. 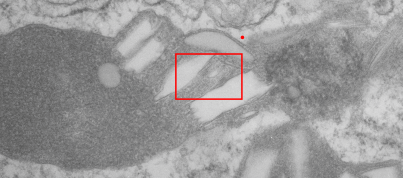
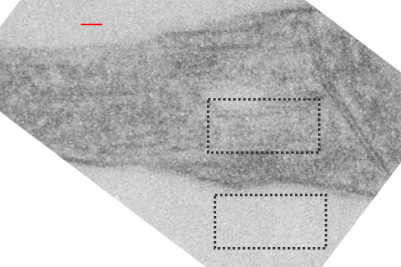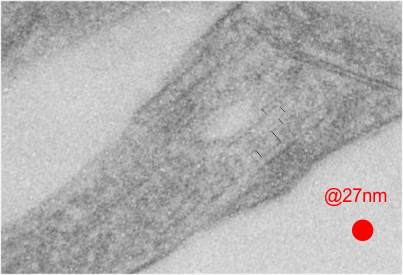
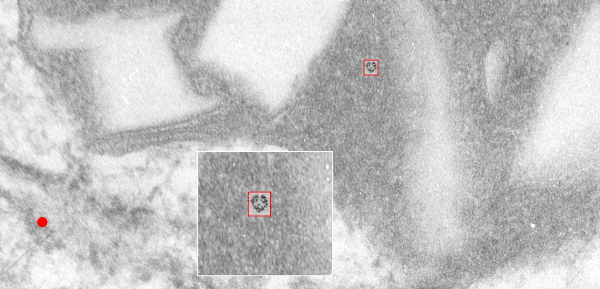
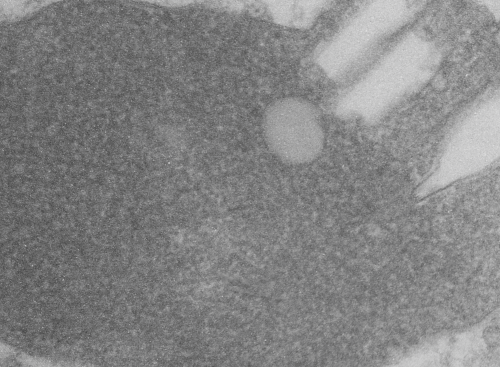
 rotated image for easier measuring and comparing the textured are (top box) and the basic epon texture (bottom box).
rotated image for easier measuring and comparing the textured are (top box) and the basic epon texture (bottom box).