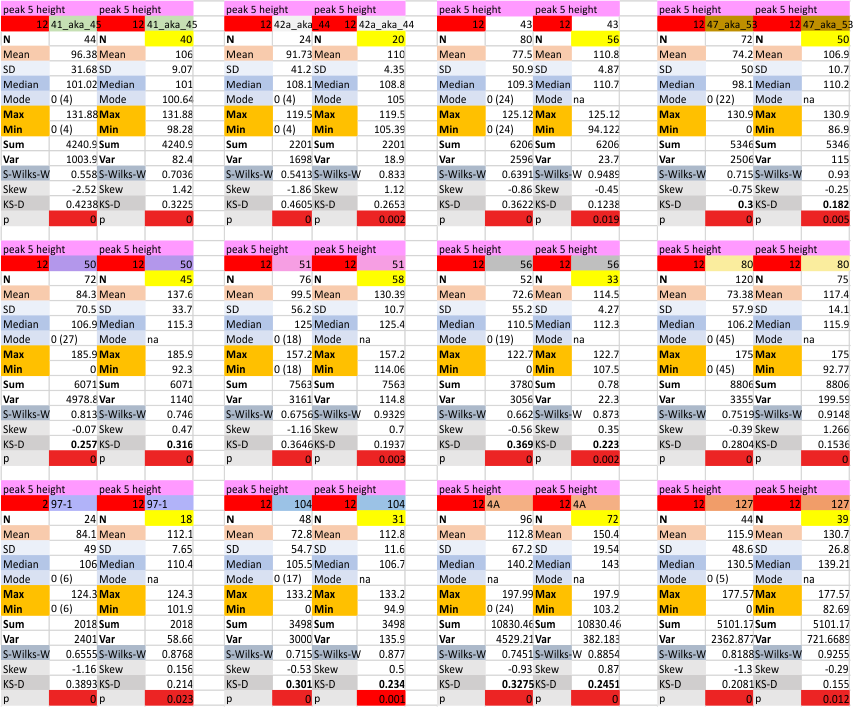
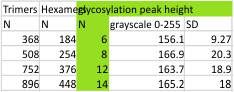
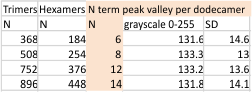
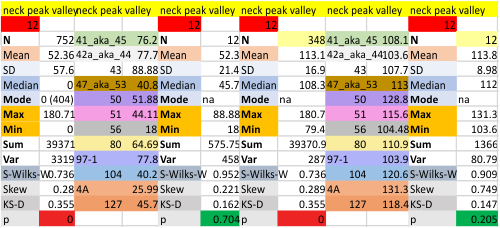
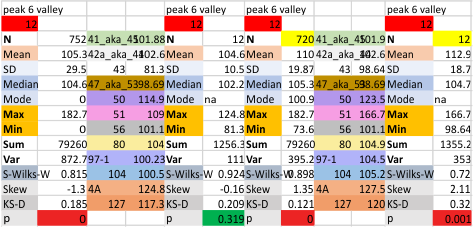
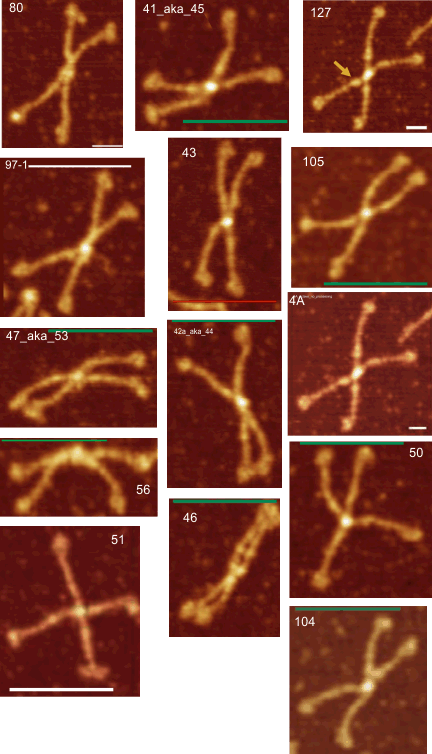
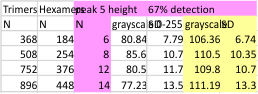
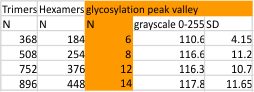
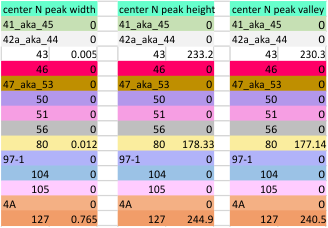
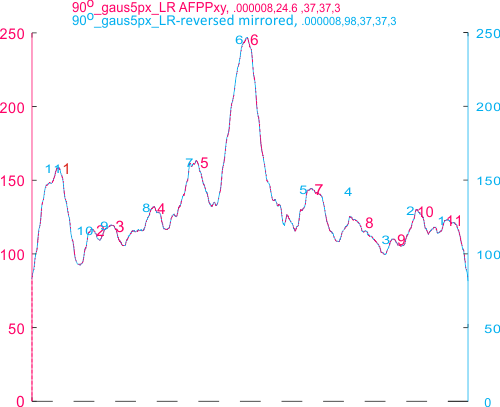
14 total dodecamers (896 trimers plotted, incrememntal addition of plots).
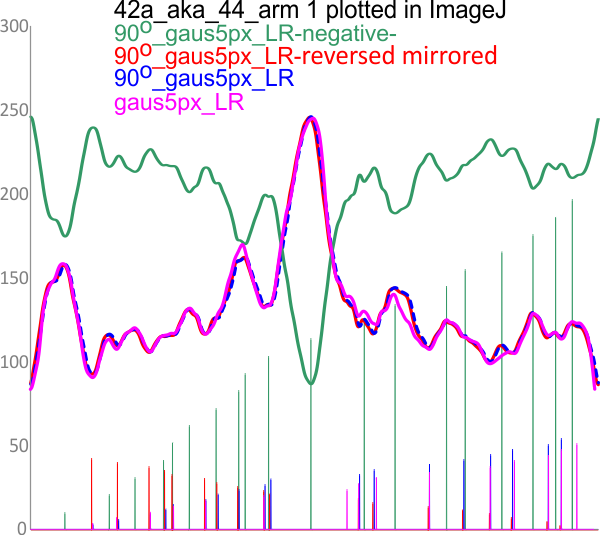
– peak widths-nm, peak height and valley-grayscale – Little changed with the signal processing, image processing filters. Plots generated in excel (the silly shoulders that excel creates that I dont know how to get rid of in excel were removed in corelDRAW by deleting those nodes on either side of the peaks).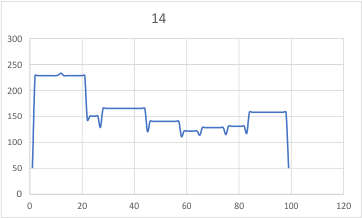
The plots are virtually identical, 8 peaks, N term peak here is NOT divided in half for each trimer but is measured as a whole peak.
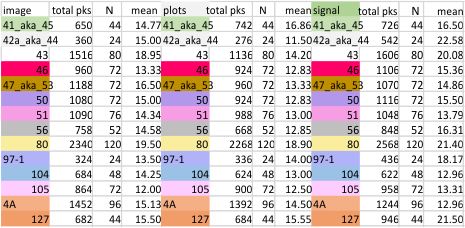
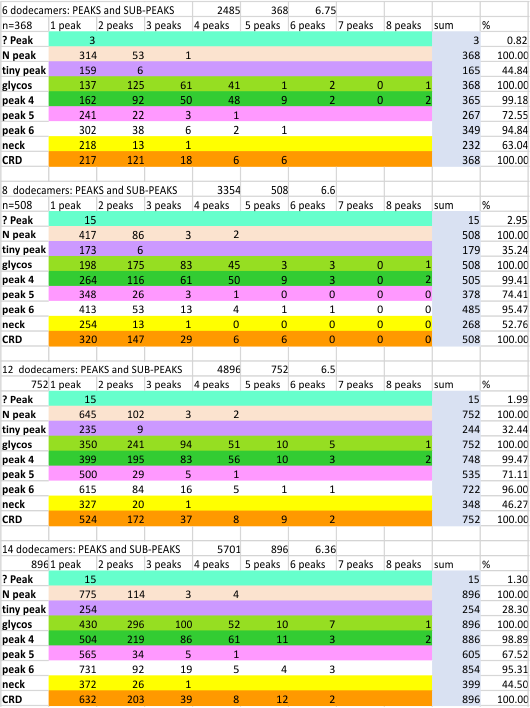
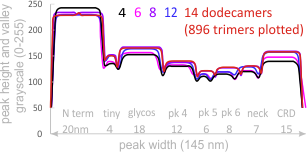
Individual plots from analyzing 4, 6, 8, 12 and 14 dodecamers are shown at the same width (@145nm) and grayscale (0-255) (below). The very infrequently detected very tiny blip present in the N term peak is not counted as one of the 15 total (8 per trimer) peaks.
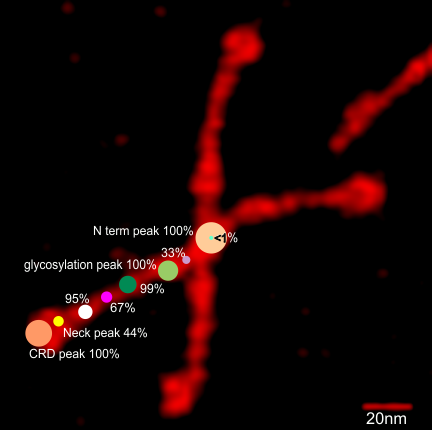
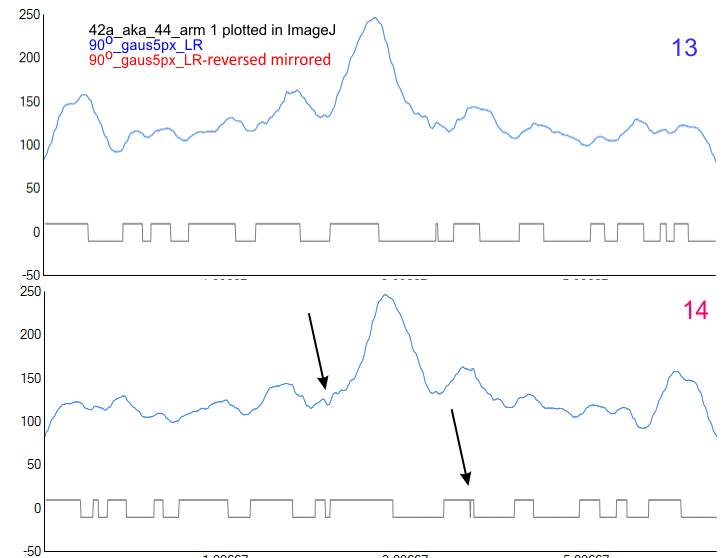
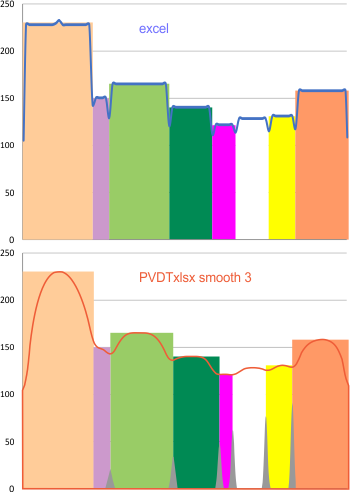
 Using the original excel plot (which has the lumpy corners) cut and pasted into the PeakValleyDetectionTemplate (using “smooth 3”) one can compare the peak detection. The tiny peak (shown in purple – and detected about 30% of the time overall) but is still visible using the PeakValleyDetectionTemplate (bottom graph). In the excel plot of 14 dodecamers (top graph) shows it clearly (tiny purple peak, on the downslope of the N term peak). Gray spikes on the baseline of the PVDT shows the detection of valleys (of the peaks) are using PVDTxlsx smooth 3. The “tiny peak” is still present, as a very tiny change in the downslope of the plot. Legend: Peach color=N terminal peak 100% occurrence (the center N peak is not shown); purple = as yet undefined tiny peak, 31% detection; medium green= glycosylation peak, 100% detection; dark green= as yet undefined peak 4, 98.88% detection; pink = narrow small as yet undefined peak 5, 67% detection; white= broad but low peak as yet undefined peak 6, 95% detection; Yellow=neck peak, 44.5% detection; dark orange= CRD peak, 100% detection.
Using the original excel plot (which has the lumpy corners) cut and pasted into the PeakValleyDetectionTemplate (using “smooth 3”) one can compare the peak detection. The tiny peak (shown in purple – and detected about 30% of the time overall) but is still visible using the PeakValleyDetectionTemplate (bottom graph). In the excel plot of 14 dodecamers (top graph) shows it clearly (tiny purple peak, on the downslope of the N term peak). Gray spikes on the baseline of the PVDT shows the detection of valleys (of the peaks) are using PVDTxlsx smooth 3. The “tiny peak” is still present, as a very tiny change in the downslope of the plot. Legend: Peach color=N terminal peak 100% occurrence (the center N peak is not shown); purple = as yet undefined tiny peak, 31% detection; medium green= glycosylation peak, 100% detection; dark green= as yet undefined peak 4, 98.88% detection; pink = narrow small as yet undefined peak 5, 67% detection; white= broad but low peak as yet undefined peak 6, 95% detection; Yellow=neck peak, 44.5% detection; dark orange= CRD peak, 100% detection.
 It seems very likely that the addition of more plots will make little difference in the number of peaks found per hexamer (15) and the relative width and height of those peaks. These images were all obtained using rhSP-D with known glycosylation.
It seems very likely that the addition of more plots will make little difference in the number of peaks found per hexamer (15) and the relative width and height of those peaks. These images were all obtained using rhSP-D with known glycosylation.
Number of glycosylations per trimer is not defined (to my knowledge) thus differences in peak height and width of the glycosylated peak could vary.
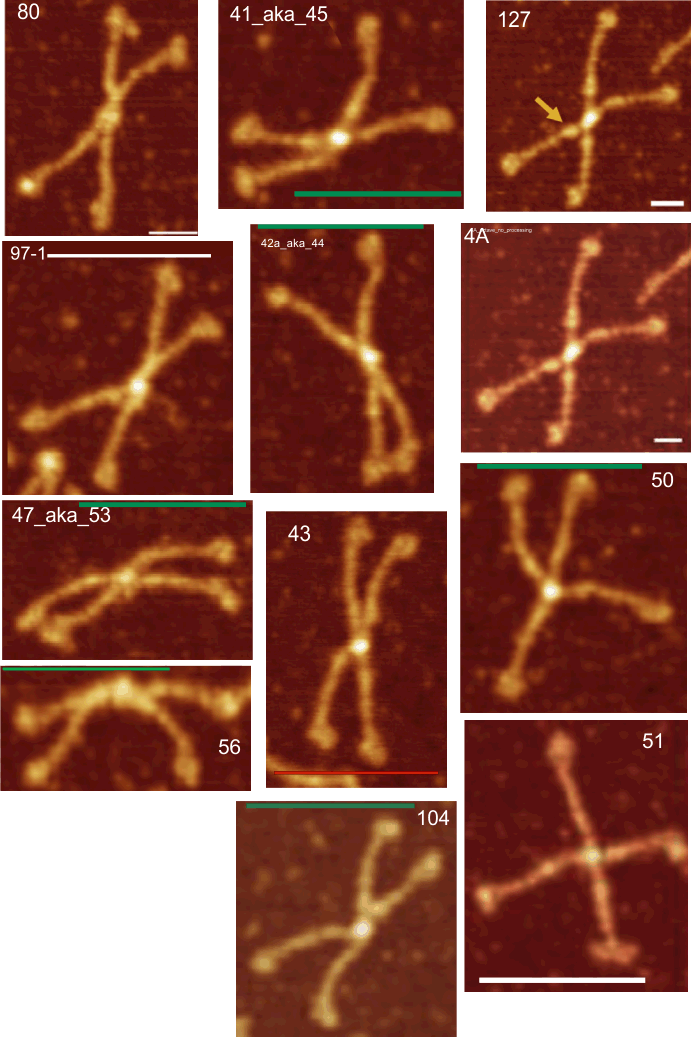
The N and CRD peaks are very consistent in relative height and width. The neck peak is often not detected – because of the variable position of the three CRD in each trimer, and that they apparently can completely obscure the neck peak during preparation by falling over it.
Over 1000 different plots of trimers comprise these figures.
Comparisons with other SP-D image (those without glycosylation, those from other species) would be valuable in helping to create a full length model of the structure of SP-D hexamers, dodecamers, and multimers.