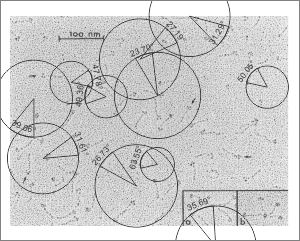
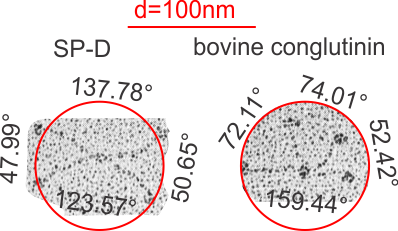
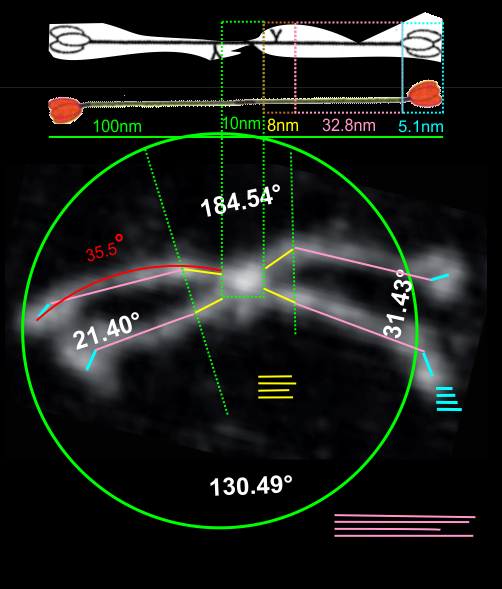
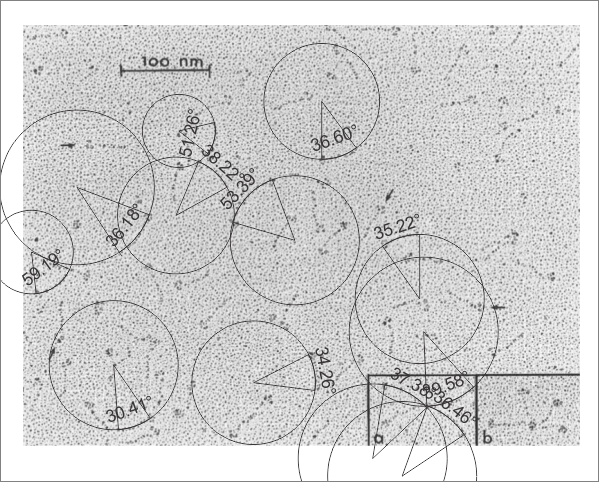
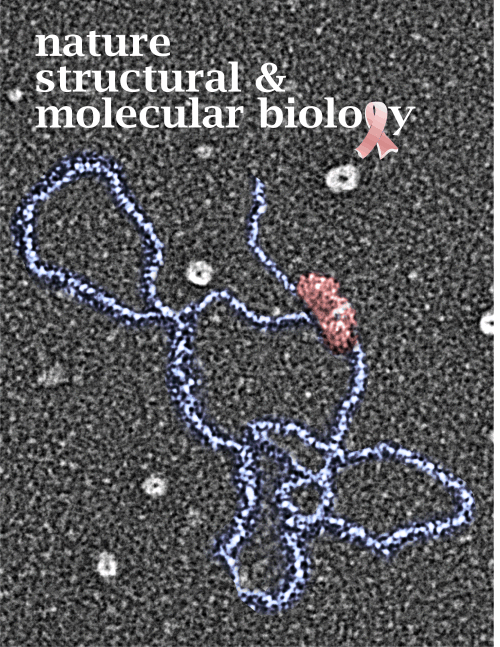
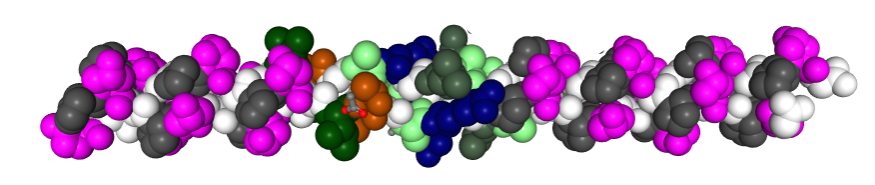
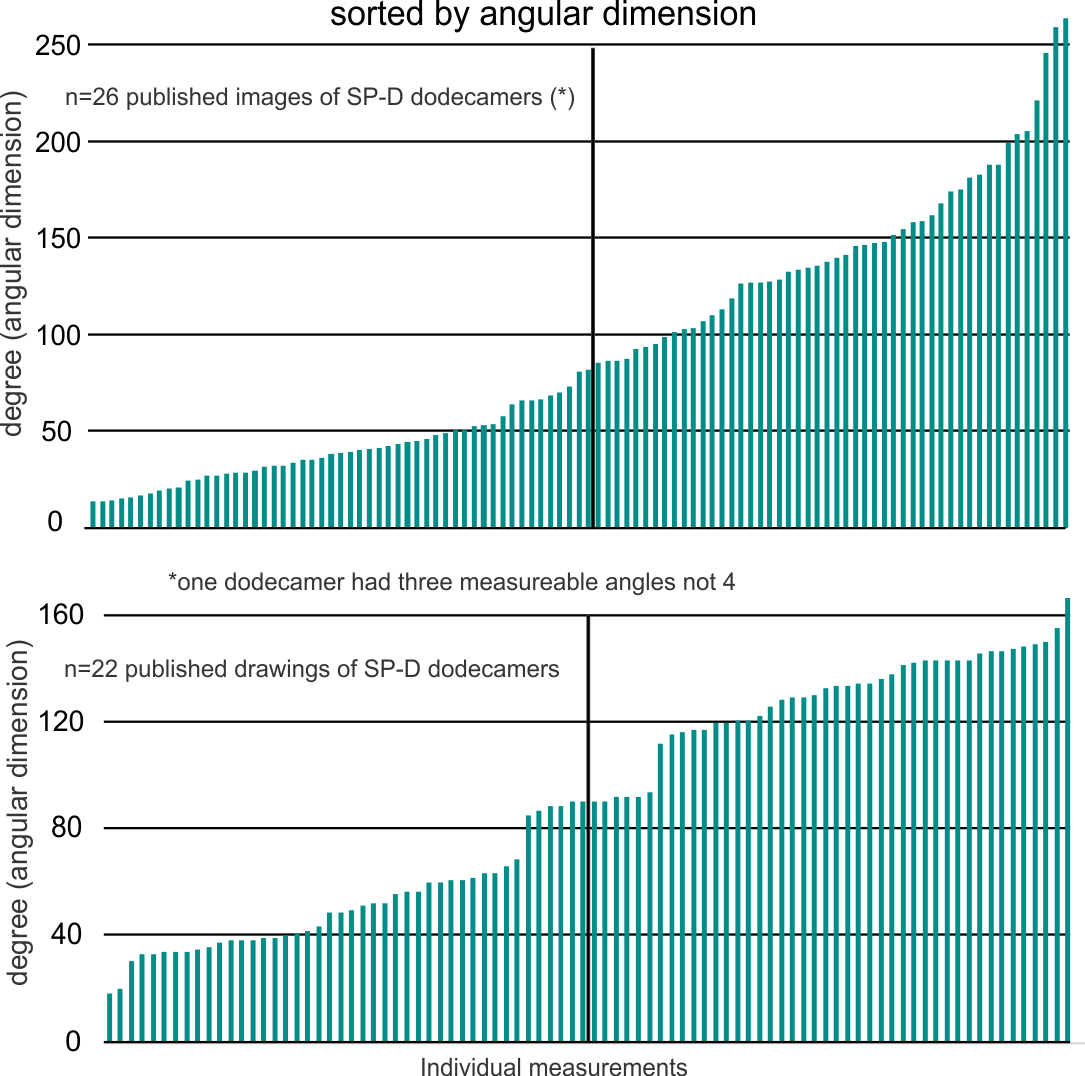
Measuring 30 SP-D dodecamers derived from numerous publications using several different methods for visualization, led me to identify this particular image as the quintessential SP-D dodecamer. The measurements then, from this particular image (most likely from the many images produced by Crouch et al, but which I have also upped the ppi and contrast on) had the closest arc’s angles in each of the individual trimer arms comprising the dodecamer to that which I measured from around 120 trimers. That is…right around 35 degrees. The distance to what looks like a bend and a bright spot in each of the trimeric arms in the electron micrograph likely represent the glycosylation site which was identified by several different investigators. This point gives the trimers the look of an arc. The carbohydrate recognition domains as they attach to the “neck” also like an angle (not yet measured). The center “bright area” of the dodecamer area where all the trimers are bound at the N terminal has a width of about 10nm, at least as is measured in this quintessential image (A collective measure the width and length of the central core of the SP-D dodecamer needs to be done). The the mean for the acute and obtuse angles of the four arms (about 30-50o and 120-140o — in this case the angles are 21.4 and 31.43 and the obtuse angles are 184.54 and 130.49 and so are a not exactly representative but close). 99% confidence level is between 37.00 and – 58.39o for the two acute center angles.
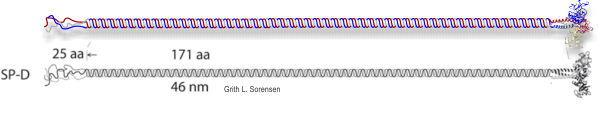
Two commonly reproduced diagrams are included at the top of the real SP-D for comparison. It is obvious that the flexibility of the SP-D dodecamer is clearly missed. And in support of the arms of the dodecamer NOT being stiff, but flexible, authors have suggested that mutant SP-D molecules (shortened collagen-like domains, deleted collagen-like domains, etc) can retain some functions in innate immunity, though sometimes differently than the native SP-D.
The purpose for making the measurements is to determine whether the fuzzyball structures made of dodecamers joined in the center at have the straight arms as depicted in nearly every single SP-D diagram.