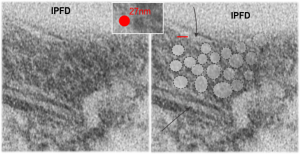
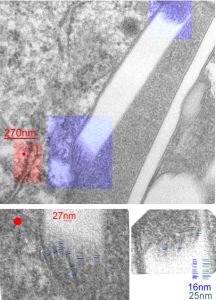
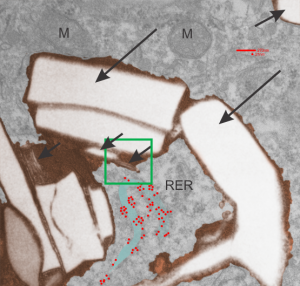
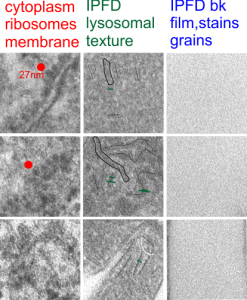
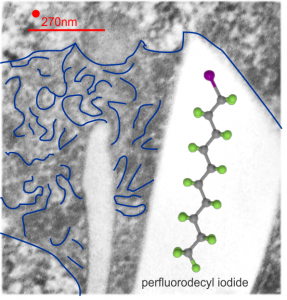
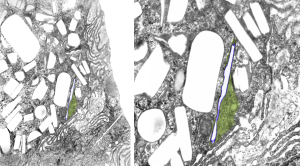
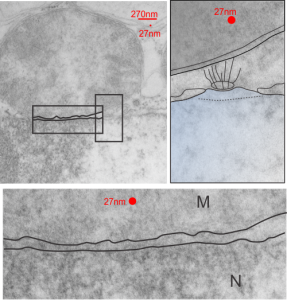
Perfluorodecyl iodide crystal periodicities are something that certainly looks like a phenomenon visible with electron microscopy which sampled such crystals after two hundred plus days residence in mouse macrophages. The crystals vary in size several orders of magnitude both in width and length, some appearing like little fractures of the whole.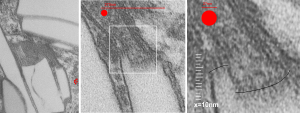 On the long ends of the IPFD crystals (within phagolysosomes, and surrounded by lysosomal enzymes which show their own type of periodicity (kind of a tubular or rounded structure — maybe both) protein strands trail out in a very distinct sweeping pattern. I have measured 15 distances between dense areas in this pattern (marked by black lines in the figure on the right and shown in array as white lines stacked on the left of that same micrograph). Taken back to the mean size of a ribosome (see image on left hand size – red dots are over ribosomes, and equate to 27nm diameter) the space between these strands coming from the long ends of IPFD crystals are about 10nm in width. I had sort of expected them to be closer to the 15nm measured in previous posts, but this is pretty close, and there is always variability from tangential sectioning and other artifacts.
On the long ends of the IPFD crystals (within phagolysosomes, and surrounded by lysosomal enzymes which show their own type of periodicity (kind of a tubular or rounded structure — maybe both) protein strands trail out in a very distinct sweeping pattern. I have measured 15 distances between dense areas in this pattern (marked by black lines in the figure on the right and shown in array as white lines stacked on the left of that same micrograph). Taken back to the mean size of a ribosome (see image on left hand size – red dots are over ribosomes, and equate to 27nm diameter) the space between these strands coming from the long ends of IPFD crystals are about 10nm in width. I had sort of expected them to be closer to the 15nm measured in previous posts, but this is pretty close, and there is always variability from tangential sectioning and other artifacts.
Speaking to artifacts…. the periodicity here cannot be attributed to some random precipitation of whatever-proteins…. since a clear and even density in background glumping of fixative and lose proteins around during fixation is seen in the “spaces” once occupied by perfluorodecyl iodide. Comparing with that background… this is pretty clear patterning.
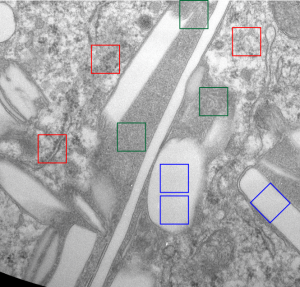
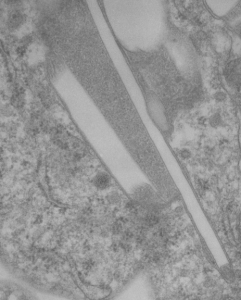
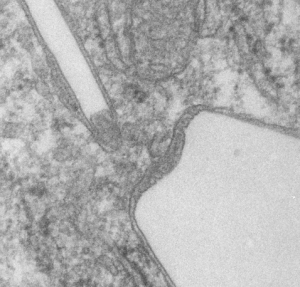
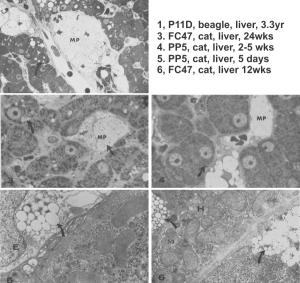
Left hand image: IPFD inclusions in a macrophage taken from mouse liver where the animal had received IPFD 267 days prior to euthanasia (details in earlier posts). This is a small portion of a single macrophage and the light areas are the footprints of IPFD crystals. Most of this image, too, is within a single variably-shaped phagolysosome, filled with IPFD crystals and a curley but tubular shaped (oligomerized?) electron dense lysosomal protein(s). Middle images is magnified from box in left hand image, and right hand image is magnified from box in middle image. Middle and right hand images show the periodicity at IPFD crystal “ends”. I am trying to find any information on the crystalline structure of IPFD from the chemical databases… but have not found it yet.