Desmosomal-mitochondrial tethering – more support for about one cadherin dimer per 13-14nm in a transmission electron micrograph fro mouse liver. 6094_15034_mouse liver 70d control for infusion of perfluorochemical emulsions. July 10 1980.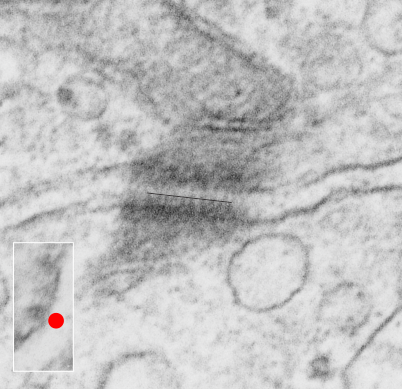
Category Archives: Electron micrographs of liver
Desmosome-mitochondrial tethering: just what I see
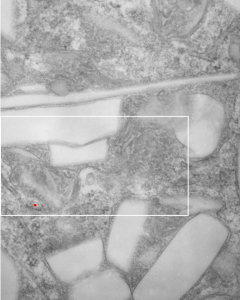
I found this particular electron micrograph which has two mitochondria and a desmosome in pretty good orientation to look at any periodicities or patterning in the substructure. It seemed to me that the central line of the desmosome, and the cadherin molecules which create it and the links to the plasmalemma (which have been modeled with molecular models) could fit best as a “sprint” type association. Those of you who are old enough to remember the plastic hair combs that looked like wishbones arranged in parallel that one could use to sort of tie up a pony tail, will recognize this flexible structure. I found, or should say “think i found” a similar type of density in the cadherin molecules in the center of this desmosome (diagrammed as that side by side “wish bone” array. Interestingly, cadherins were mentioned to one group to come in a dimer which would very easily form this springy wish bone structure…. and listed the “half wish bone” as one option as a molecular model of the desmosomal cadherins. One of those articles has a great 3D image which doesn’t look exactly like what i see, but is close found here – and another is this title Studer, Daniel & M Humbel, Bruno & Chiquet, Matthias. (2008). Electron microscopy of high pressure frozen samples: Bridging the gap between cellular ultrastructure and atomic resolution. Histochemistry and cell biology. 130. 877-89. 10.1007/s00418-008-0500-1.
. I actually don’t like the idea of calling the arrangement “untangling desmosomal junctions with knots” as i think if they were knots….there would be no perfectly wonderful order that can easily be seen with ordinary transmission electron microscopy. It is also very appealing to have flexibility within these junctions as the wishbone (wish bone) alignment would afford…. that would be just really fun. 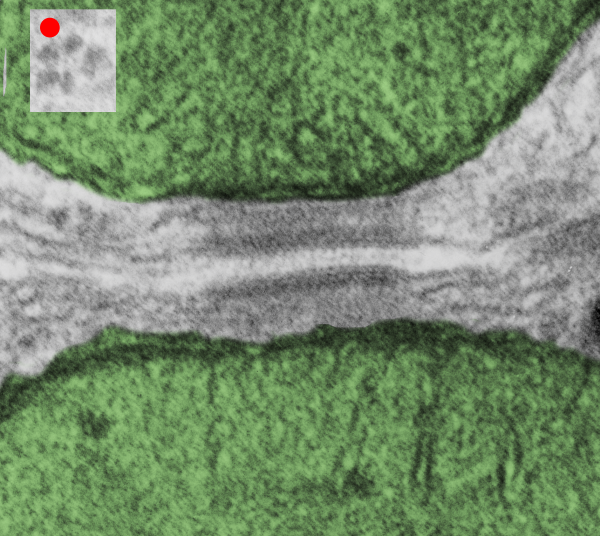 So here is the same electron micrograph as yesterdays post but the mitochondria are actually fuzzed green in a separate layer. Red dot=a ribosome, approximately 27 nm in diameter. The absolutely regular arrangement of cadherins is seen in the middle of the desmosome…. no knots, no tangles, no mess…. just regularly spaced. This micrograph is not retouched to emphasize anything…. it is just the way it looks on the negative. The plasmalemma of the two cells is not in exact cross section and so it is fuzzed. The span between densities both on the plasmalemmal sides and the center densities of this desmosome work out to be about 1/13-14nm. I have not put that together with any models yet and in the image below just sized the molecular structure according to the TEM, not using the actual size (just using the shape).
So here is the same electron micrograph as yesterdays post but the mitochondria are actually fuzzed green in a separate layer. Red dot=a ribosome, approximately 27 nm in diameter. The absolutely regular arrangement of cadherins is seen in the middle of the desmosome…. no knots, no tangles, no mess…. just regularly spaced. This micrograph is not retouched to emphasize anything…. it is just the way it looks on the negative. The plasmalemma of the two cells is not in exact cross section and so it is fuzzed. The span between densities both on the plasmalemmal sides and the center densities of this desmosome work out to be about 1/13-14nm. I have not put that together with any models yet and in the image below just sized the molecular structure according to the TEM, not using the actual size (just using the shape).
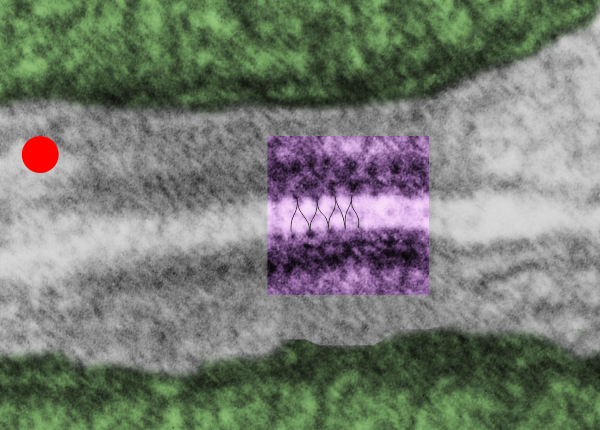

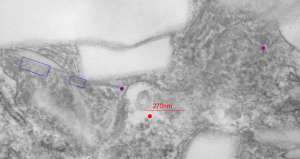
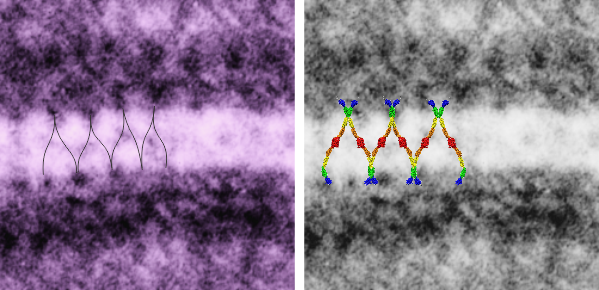
So in the second figure you can see an area that i enlarged, cut and pasted in photoshop, and enhanced in contrast and colored purple and ADDED what i think are the lines of the wishbone arrangements of the cadherins. Densities (increased in contrast here) work out to about 1 for every 13-14nm.
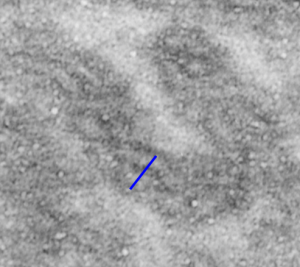
Into two enlargements (one the purple box above and another with the molecular model of a cadherin dimer, copied and mirrorred, fit really nicely into what might be a flexible, stretchy springy type portion of the desmosome. Just a thought…. ??
6110_5080_mouse_f_liver_36,200x_4x
Cadherin molecules overlying a desomosome – mitochondrial tethering
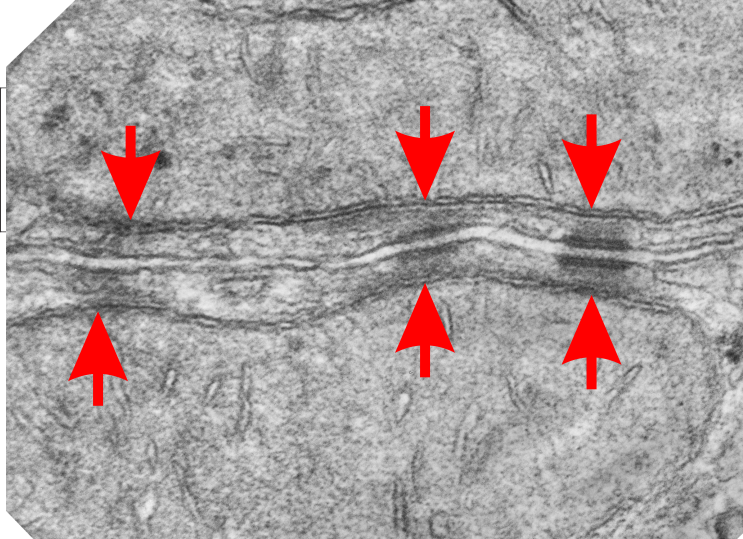
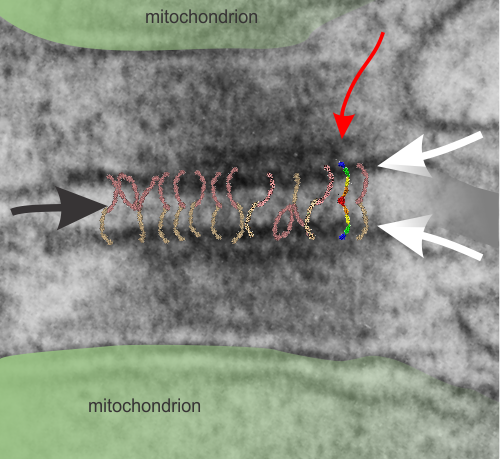
I don’t know much about desmosomes, but they do make unique junctions with nuclear pore intermediate filaments and also very definite connections with mitochondria. The mitochondria – intermediate filament connections with desmosomes show a couple nice ultrastructural changes from the routine: 1) the mitochondrial outer membrane is flattened, or straightened in the area of connection with the intermediate filaments over the desmosome, and it is also a little darker. and 2) the mitochondrial body itself is drawn towards (at least that is the way it looks… as if there was a dragging force) the desmosomal intermediate filaments. The central line of the desmosome is not always visible when i see these tethers (often with two mitochondria in adjacent cells and a single desmosome, but there are molecular models of the cadherins (several) that I cut pasted masked and reduced to fit in the appropriate intercellular space where they would appear. It is a good lesson is relative “size” and “shape”.
Micrograph: Pale green semitransparent mask is over two mitochondria, one each in two adjacent cells. Red arrow points to the inner dense layer (maybe i will add the known proteins there “in scale), the white arrows point to the plasmalemma of the two separate cells, the black arrow points to the center dense “knotted” region, (maybe i will measure distances between visible knots) and the molecules (several taken from the internet, look similar and were pasted into the intercellular region (see demarcation from right hand side transparent grey.
Triple desmosomal mitochondrial tethers
Both sides of two adjacent hepatocytes show two mitochondria each linking to three desmosomes. Nice to see such symmetry and the implications are quite amazing, in that whatever is happening at the plasmalemmae of these cells is happening “together”, really nice intracellular communication. Red arrows point to the inner mitochondrial membrane, just above where the intermediate filaments of the desmosomes approach the outer mitochondrial membrane. I wish i could say that I saw some cross sections of intermediate filaments here, but i didn’t. The double desmosomal mitochondrial junction on the left is more tangential than the one on the right. The pair in the middle is intermediate.
Wedding cake crystals: perfluorodecyl iodide
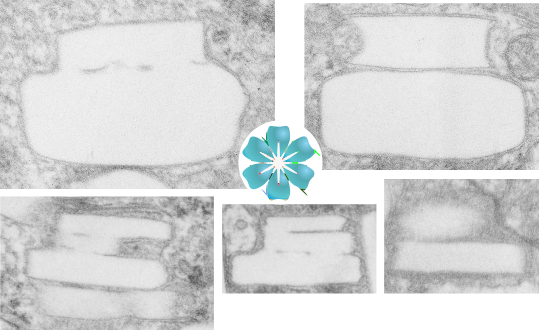
These crystals are certainly odd. I have for many years wondered at their layered cake appearance, and today just found it so funny that i had to make a spoof about it. So here are a few images taken from within a single hepatocyte phagocytic cell (like a resident macrophage) and these layered cakes ‘so to speak’ are oriented with the largest layer at the base and stacked vertically. The crystals appear to be stacked round layers, but probably not always, maybe with a chance orientation of smaller layers “up one”, I have not done any counting to see whether they might be “reducing in size as they stack”. The rounded bottom layer (not shown in these images) is seen in cross section sometimes as a full circle, other times as a tangential cut with one straighter edge and one rounded edge, just as one would find in a tangential cut of a layer cake.
At any rate, these are clues about the structure of perfluorodecyl iodide crystals in vivo.
Perfluorodecyl iodide crystals in vivo: big
Just reviewing some slides of liver of mice given infusions of an emulsion of perfluorodecyl iodide…. some of these crystals which completely fill to overflowing hepatocyte macrophages (multinucleated macrophages or macrophages fused into multinucleated giant cells following the foreign body response to IPFD infusion). Just saying here, some of the IPFD crystals are upwards of 30 microns long, and vary in width from 10nm to many microns. There is a very obvious directionality, always longer than wide. Iin terms of the actual molecular polymerization of IPFD, which is necessary to establish what is “wide” and what is “long” — that remains to be determined. But for now, these macrophages are about 90% “other” with half dozen nuclei in a localized cytoplasmic – crystal free zone together.
And regarding spleen, it has far fewer cells containing IPFD than I would have expected (just a relatively small number of reticuloendothelial cells show collections of crystals (compared to liver from the same animal which does have multinucleated cells full of crystals). So that is kind of interesting.
55 days after 100cc/kg of a 10% IPFD 5%F68 emulsion, the hepatocytes themselves are very well protected from inclusions of IPFD…. there are some probably, but not many. Hepatocytes themselves look nice, but the clusters of macrophages, without eliciting much of a fibrous tissue response are neatly circumscribed.
If…Then
I sometimes get the uneasy feeling that if i see a pattern in an electron micrograph that someone is going to say, ah thats just fixation artifact… and of course they are right. All testing is “fixation artifact” but its the purview of each observer to determine what is background noise and what is real structure.
In the case of looking at perfluorochemicals left behind in tissues, consensus says that there is “nothing” in that empty footprint, just the embedding medium (in this case EPON 812). So there is a texture to the embedding medium, no question, but nothing I can perceive, and so easily compared to the texture of surrounding tissues, so that when I see a pattern in the tissue I am pretty convinced that it is the presence of the proteins . (They are influenced by the fixative and type and length of dehydration and other stuff not excluding the warping and shifting under the electron microscope. All those factors play a role). It is one of the reasons that an internal control for size (and i am suggesting background texture) in electron microscopy is really important. I CAN list a short list of variables to be aware of (my knowledge is just observational) which begin even before biopsy, including the time of the last meal, 1) circadian cycle, 2) health of the tissue (in vivo or in vitro –doesn’t matter) 3) mode of analgesia or anesthesia at biopsy (paralytics – solutions with ions (as in perfusion) 4) time till tissue gets into fixative, 5) size of the tissue block, 6) type of fixative, 7) length of time in fixative, 8) length of time and chemicals used for dehydration, 9)type of plastic, 10) section thickness, 11) staining compounds, 12) TEM specs, KV, etc…. so I am willing to accept that not all parameters are under one’s control.
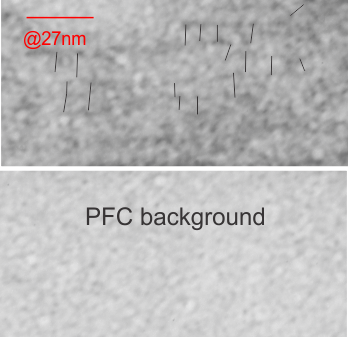
I will however say that the pattern below (in a portion of a macrophage in the liver of a mouse given 10%IPFD in 5%F68, 267 days post infusion) shows a linearity I would not be willing to ignore. low mag micrograph top, for orientation, next below is micrograph of the red box area which shows the periodicity. That is just one of the linear arrangements that are visible in these tissues. 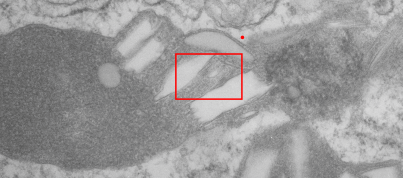
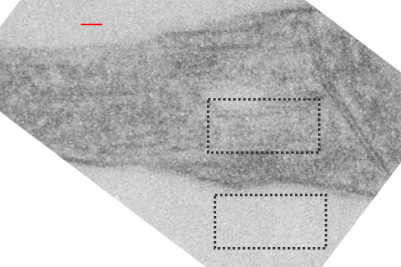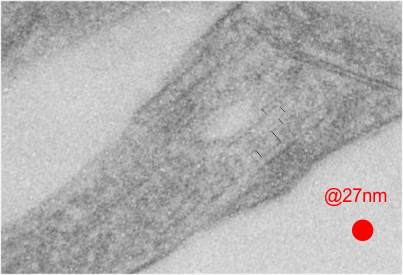
 rotated image for easier measuring and comparing the textured are (top box) and the basic epon texture (bottom box).
rotated image for easier measuring and comparing the textured are (top box) and the basic epon texture (bottom box).
Phagolysosomal and crystal patterns in IPFD inclusions in liver
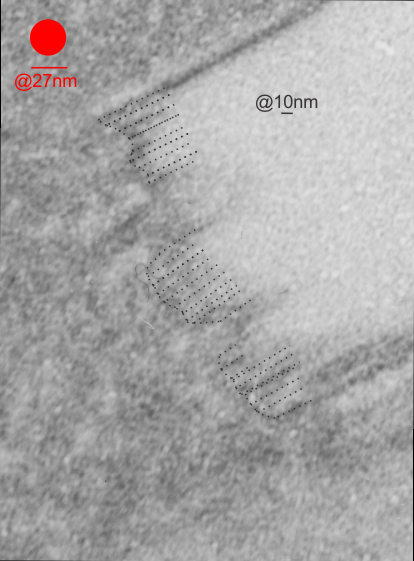
Two prominent areas of patterning are apparent in phagolysosomes in liver after IPFD infusion. The IPFD crystal itself, and at least four distinct protein patterns in the phagolysosomal proteins. The first image below is an enlargement of the red rectangle, and I have marked off with lines some of the pattern at the long end of the perfluorodecyl iodide crystals which seemed to work out to about 10nm in the short dimension. 23 measurements made at the end are marked off with dotted lines. A red dot=the dimension of a ribosome taken at the same magnification for reference. Below that is the low magnification image (unretouched — I removed a scratch from the top image but no data were altered as it was not in the crystal itself). Black boxes correspond to four very distinct protein organizational patterns within a single phagolysosomal body which also included the crystal in the top image. I am pretty confident that these patterns are a clue as to which lysosomes are present, and 4 may not be the only ones which show oligomerization into ultrastructural patterns. One thing to think about is that the kupffer cells (likely) in liver acting as macrophages which engulf (and hold onto for months) IPFD crystals will have a different panel of enzymes for lysosomes (and peroxisomes) than hepatocytes, and seemingly can become multinucleated under the conditions of massive IPFD inclusions.
Perfluorodecyl iodide: a peroxisome proliferator
Going through archived images of the livers of mice which received infusions of iodo containing perfluorochemicals, I ran onto a hand-written comment which made me laugh. It was “funky print given under pressure to LC Clark (Jr) that perfluorodecyl iodide increases tremendously the number and size of microbodies (aka peroxisomes) in the mouse liver” (words in italics added today for clarity). Apparently no one wanted to hear that news in 1973 or 4… perhaps because it was a “damning” mark on the possible use of perfluorochemical formulations in general, and specifically for artificial – blood substitutes.
Wikipedia summary is here: “They are involved in catabolism of very long chain fatty acids, branched chain fatty acids, D-amino acids, and polyamines, reduction of reactive oxygen species – specifically hydrogen peroxide; – and biosynthesis of plasmalogens, such as ether phospholipids critical for the normal function of mammalian brains and lungs. They also contain approximately 10% of the total activity of two enzymes in the pentose phosphate pathway, which is important for energy metabolism.”
Below are three images, one of liver which shows the very large peroxisomes, in this particular micrograph (not all shown here) they were a volume density of about 30% of the cytoplasm which puts IPFD up there with the best of the proliferators; mitochondria for comparison were about 33% of the cytoplasm. Consensus has peroxisomes at more like 2% of the volume density of the hepatocyte under unstimulated conditions.
The story will be complicated as there are more than 30 enzymes which are in peroxisomes: and the patterning that is seen in the enzymes within the phagolysosomes containing IPFD looks to be similarly wound as the enzymes within peroxisomes in the hepatocytes (top two image groups). From the internet the middle three images (used without permission as a screen print of the insanely egotistical Dr. Jastro TEM collection who most certainly used public funds for this research and collection of micrographs and yet thinks he has ownership of these images…. not civil at all in my opinion and shame on him) is an hepatocyte peroxisome (which i measured against a nearby ribosome and also the micron bar marker, for size) shows a coarse texture, somewhat coiled. The latter and my peroxisome images (top), added from a portion of an hepatocyte from the liver which received IPFD and measured against a nearby ribosome, show similar “oligomerization” or “substructural” protein patterning.
Some protein(s) is creating this texture. It would be interesting to know if the protein in these microbodies (aka peroxisomes) is the same as that which is found in abundance in the phagolysosomes containing IPFD. Bottom micrograph is a portion of hepatocyte cytoplasm from IPFD infused mouse more than 200 days post infusion from which my (top two) images are derived.


Endosomes in phagocytes containing perfluorodecyl iodide crystals show unique substructure
The phagolysosomes in phagocytes which containing perfluorodecyl iodide crystals (in this case the perfluorocarbon has been retained in the body for more than 250 days – mouse liver) show huge amounts of lysosomal enzymes. The collection of macrophages (even multinucleate macrophages) don’t appear to initiate a huge collagen response, and inflammatory cells such as neutrophils, just are not that common, even when phagocytes are plentiful.
The ultrastructural characteristics of the lysosomal enzymes contents within phagolysosomes containing IPFD crystals can vary dramatically within the same cell and in adjacent lysosomes, and within the same lysosome. This varies from a very fine homogeneous non-textured content in some areas to areas with a more coarse fibril-like texture, appearing like a lumpy (sausage link type) strand or a twisted strand (of two separate elements, or a two strand braid). In some cases the fibrils appear to be solid on cross section, but in other cases more like a tubule. This underscores the variability that would be found in a twisted fibril, allowing for the explanation of both “hollow” and “solid” profiles on cross section.
Transition zones between the types of enzyme ultrastructure are found within the same lysosome suggesting some protein overlap or continuity. There are some areas of very dense accumulation of proteins.
IPFD crystals may be large or small, and most display some obvious directionality, usually the longer side having a thin dense protein (or trilaminar membrane?) coat, but at the ends showing considerable “striping” or dragging out, or whispy periodicity.
IPFD crystals often appear in cross section to be more rounded, or even round.
Some common characteristics of the fibrillar texture of lysosomal contents of IPFD filled lysosomes follows:
- relatively even diameter at widest and narrowest areas
- often looped and tangled, folded looking
- equally even spacing seen as a predominant lucent area surrounding each strand
- often running in parallel
- higher mag shows sub-layering
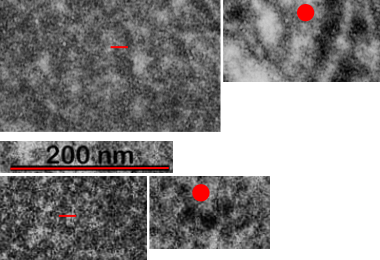
This image below shows a very prominent (yet small in terms of nm) level of periodicity. May not be the smallest since I havn’t be able to find any information on the actual crystal chemical structure of perfluorodecyl iodide), but pretty small for conventional electron microscopy to propose, but this is so obvious that it certainly warrants comment. The single space (inset is same size as lower image and placed on an identical image, enlarged so that the lines (periodicity) could be seen. The span is about 9-10nm line to line, and the fiber arrangement is double that…. between 19-20nm. It appears that the protein(s) oligomerized here can come as an interconnected (maybe twisted) double strand.
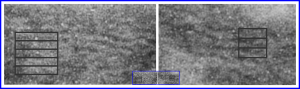

Lower magnification image of the portion of a macrophage from which the inset to the right and above (white box) and the measurements of distance between fibrills (white lines)and smallest periodicity (blue box) measurements were made. Red dots=ribosomes at an estimated 27nm for reference; blue line = smallest periodicity; purple line = the 9-10nm pair, twisted, representing the fibrills (about 20nm in thickness if section is longitudinal, or diameter ir fibrill is cross sectioned. A measurement of about 20+nm is found from center of one fibrill, perpendicularly, to the center of the adjacent fibrill, (white connected lines on the bottom micrograph show distances between fibrils measured. Image at bottom has blue line of @20+nm –the approximate diameter of the “fibril”, and the dividing line of the two strands is visible.