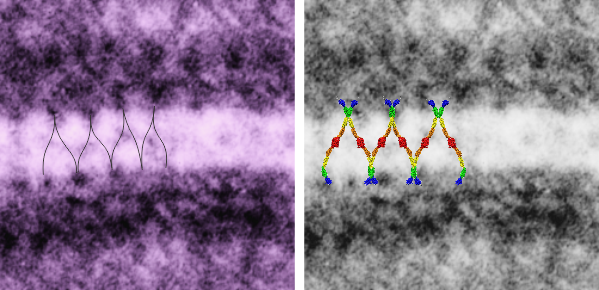
There are several nice things about this micrograph of a desmosomal mitochondrial tether (i.e. tethered via the intermediate filaments of the desmosome). One nice thing is the very rigid, and parallel membrane-leaflet quality of the plasmalemma where there is the desmosomal protein complex, and it also exhibits a very even and well defined separation of the outer and inner leafelets of the plasmalemma (at least this is what i think is shown in the micrograph below). I have marked the two leaflets on the cell membranes on the adjacent cells with black lines. There are obvious periodicities within the inner and outer membrane leaflets too… not withstanding the density of the lead citrate and uranyl acetate stains and the quality of fixation) which seems to present overall patterning in the entire micrograph. I have marked the extracellular lines (which are not likely to be random as they appear, but more likely to be organized attached wishbone shapes (mirrored vertically and copied horizontally) shown in an earlier post but the lines here have been drown over actual densities in the extracellular (intercellular) space of the desmosome. Green blobs mark periodicities within the two leaflets of the plasmalemma, and the pink blobs mark the densities which are extracellular but likely adjacent, maybe even part of (like the cadherins in desmosomes) those proteins. (green blobs are INTERcellular, the pink blobs are within the plasmalemma (transmembrane). Electron micrograph on left is unretouched, on right, contrast and brightness enhanced and vector overlays show densities which can be matched to image on the left. This stub-tail monkey hepatocyte came from an animal that received two test doses of perfluorodecalin (a very small amount) a year prior, which in my experience has nothing to do with these tethers. Shown immediately below the outer leaflet of the plasmalemma is what appears to be a regular compartmentalization (shown as rectangles) likely representing a highly structure arrangement of plakophilin and plakglobin and maybe some desmoplakin and a bit of vimentin. The number of densities (green or pink, and likely connected parts of the cadherin dimers, turn out in this micrograph to be about 9 nm… a little close than found in mouse micrographs so far. 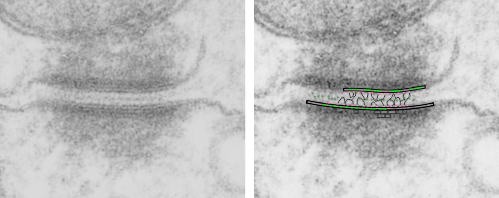
Category Archives: Desmosomal mitochondrial associations
Stub tail monkey hepatocyte desomosomal-mitochondrial tether
These tetherings between desmosomes and mitochondria are pretty plentiful. Here is another example of one such from a hepatocyte from a stub tail monkey. This one shows some very distinct banding…. a little more demarcated than liver, I wonder if this is just opportune sectioning or some molecular difference in the components. Enlargement of the desmosome below top image.
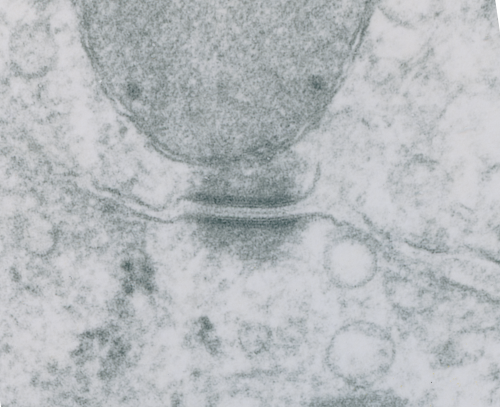
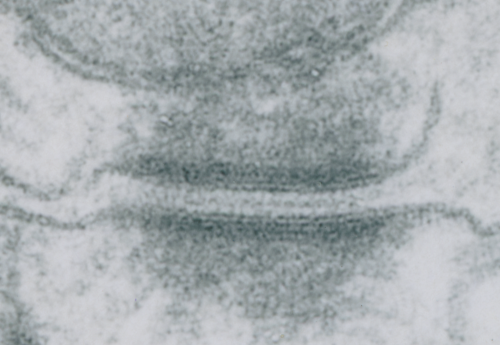 I looks in this image like the plasmalemma of each cell has a very rigid separation of the two leafelets.
I looks in this image like the plasmalemma of each cell has a very rigid separation of the two leafelets.
Desmosomes and mitochondria tethered together
Looking still for some molecular models which fit the images I see for desmosomal mitochondrial junctions (adjacent structures tethered by intermediate filaments). These junctions are so prevalent that I cannot understand why they are not part of the everyday histology lessons. This particular mitochondrion and desmosome are from a mouse liver (control for other studies) . The sectioning and orientation are no too bad though it would be nice to have even more detail (haha as much detail as one might get from electron tomography, which isnt going to happen for me so i will do the best with what i have).
The molecular models of desmosomal proteins, which are pretty widely published, may have relevance to the structures here. The first think I am pretty convinced about is the stretchy hair-pin zigzag like connection of the cadherin molecules within the center of the desomosomal structure, and the little periodicity that is apparent (10-14nm spacing gets calculated from this image). I found a nm marker only on the molecular model for intermediate filaments which was 5nm (a very helpful marker indeed) and it comes close to confirming what I have as the magnification on the micrographs as judged by a 27nm ribosome standard. The proteins overlying the TEM here are just forced to size, as there were no nm markers on those models… and the position of the one large molecule (desmoplakin) is off to the side waiting to be figured out). There are big gaps between what I see with TEM and what the molecules say…. it is fun to try to figure it out.
Upper micrograph, unretouched, also with rectangle for enlarged and pseudocolored image below. The bottom image has the intercellular space colored green, and you can see the cadherin springy wishbones above it, and also the plasmalemma and likely other proteins (plakophilin and plakoglobin) sort of carelessly put in place, and the desmocolin off to the side in an “unknown” placement. Red dots are ribosomes (for size reference).
Stub tail monkey liver: desmosomal mitochondrial tethering
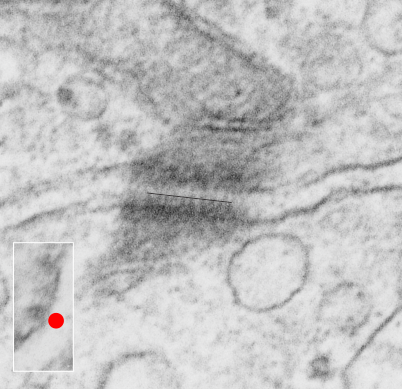
This is just a micrograph to justify the idea that the desmosomal mitochondrial tethering, or junction or des-mites as i have aptly named them (as opposed to pore-mites) for nuclear pore mitochondrial tethering, are pretty universal. Here a single tether is in a stub tail monkey hepatocyte, from tissue taken back in the 1970s while studying the effects of infusing artificial blood emulsions (perfluorochemical based blood substitutes) . This particular monkey (Maccaca speciosa, probably female) did receive two test doses of a perfuorodecaline emulsion, just a tiny amount, 0.05cc/kg of a 10% emulsion PP5ct and 5% F68) (Lee Clark Jr named all his emulsions, this was EM#750428). no perfluorochemical droplets were seen in this hepatocyte. Sac date was 5 10 76.
So desmosomal mitochondrial junctions are here. Mitochondrion is in top part of micrograph, desmosome is attached to the down pointing portion.
More pseudocolored desmosomes and mitochondria – tethered
Here is another pseudocolored desmosome with portions of two mitochondria top and bottom parts of this electron micrograph. It was difficult to determine exactly where the plasmalemma from each of these two cells went at the point of the desmosome, but I thought long and hard about which part of the structure they were. It seemed to me that there was a slight electron lucency just under the plasmalemma on each side, therefore this is the way I pseudocolored (with pink) the cell membranes. The densities within the desmosome itself looks like there are three rows… the central dotted (periodic) line where the cadherin molecules knot together (my guess is this is a totally symmetric arrangement, not at all random like suggested by some) and perhaps another periodicity (well not perhaps…. it is pretty striking) which lies a little separated from the plasmalemma. I don’t know if any of the models of the cadherins show a “lump” structure before the transmembrane part… ? That will take some searching.
Top image is unretouched transmission electron micrograph of a desmosome, as mentioned, mitochondria portion seen top and bottom. The box in this image is what is enlarged in the second image. There are two very prominent intramitochondrial granules, especially the one in the mitochondria at the bottom of the micrograph.
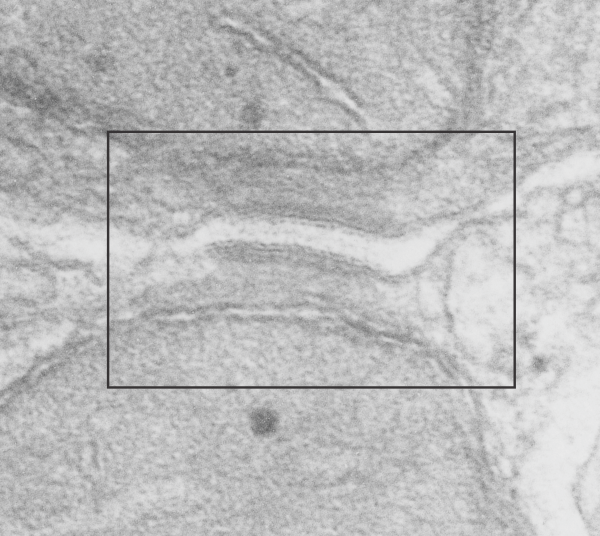 This image is from the boxed area above, thus enlarged, pink is the plasmalemma, orange is the area just under the plasmalemma of each cell and into the space of the desmosome. Blue is what I see as the possible densities of cadherin molecules.
This image is from the boxed area above, thus enlarged, pink is the plasmalemma, orange is the area just under the plasmalemma of each cell and into the space of the desmosome. Blue is what I see as the possible densities of cadherin molecules.
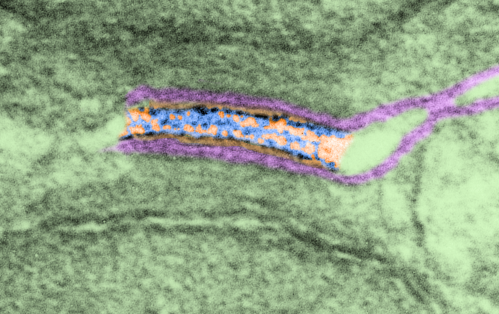 In this inset the periodicity of the outer part of the desmosome (probably still cadherin molecules) is a spacing about 1/15nm not too different from that found in a previous post at something around 1 density for each 13-14nm spacing. The periodicities on the central dense line of the desmosome in this micrograph might be something around 18nm spacing… I would have preferred if the densities came out one-to-one, but anticipate that in other assessments that it might do just that. But for now, i just count what shows up. 6102_5070_mouse_female_control.
In this inset the periodicity of the outer part of the desmosome (probably still cadherin molecules) is a spacing about 1/15nm not too different from that found in a previous post at something around 1 density for each 13-14nm spacing. The periodicities on the central dense line of the desmosome in this micrograph might be something around 18nm spacing… I would have preferred if the densities came out one-to-one, but anticipate that in other assessments that it might do just that. But for now, i just count what shows up. 6102_5070_mouse_female_control.
Desmosomal-mitochondrial tethering – more
Desmosome-mitochondrial tethering: just what I see
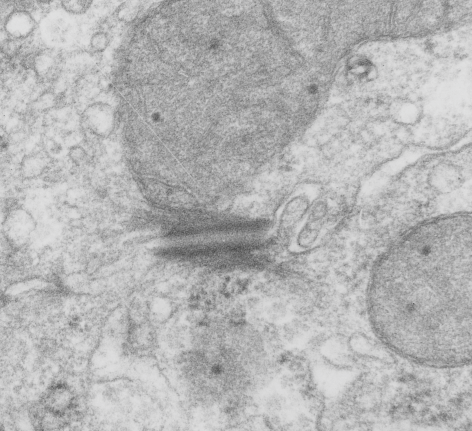
I found this particular electron micrograph which has two mitochondria and a desmosome in pretty good orientation to look at any periodicities or patterning in the substructure. It seemed to me that the central line of the desmosome, and the cadherin molecules which create it and the links to the plasmalemma (which have been modeled with molecular models) could fit best as a “sprint” type association. Those of you who are old enough to remember the plastic hair combs that looked like wishbones arranged in parallel that one could use to sort of tie up a pony tail, will recognize this flexible structure. I found, or should say “think i found” a similar type of density in the cadherin molecules in the center of this desmosome (diagrammed as that side by side “wish bone” array. Interestingly, cadherins were mentioned to one group to come in a dimer which would very easily form this springy wish bone structure…. and listed the “half wish bone” as one option as a molecular model of the desmosomal cadherins. One of those articles has a great 3D image which doesn’t look exactly like what i see, but is close found here – and another is this title Studer, Daniel & M Humbel, Bruno & Chiquet, Matthias. (2008). Electron microscopy of high pressure frozen samples: Bridging the gap between cellular ultrastructure and atomic resolution. Histochemistry and cell biology. 130. 877-89. 10.1007/s00418-008-0500-1.
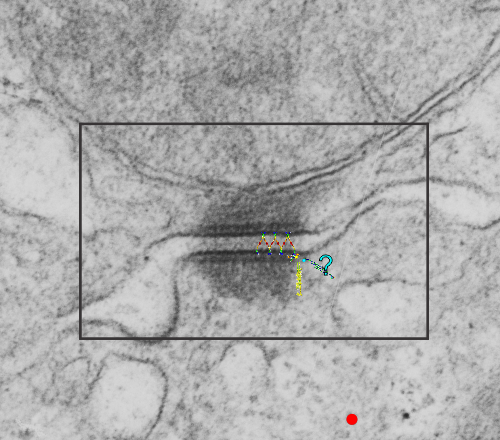
. I actually don’t like the idea of calling the arrangement “untangling desmosomal junctions with knots” as i think if they were knots….there would be no perfectly wonderful order that can easily be seen with ordinary transmission electron microscopy. It is also very appealing to have flexibility within these junctions as the wishbone (wish bone) alignment would afford…. that would be just really fun. 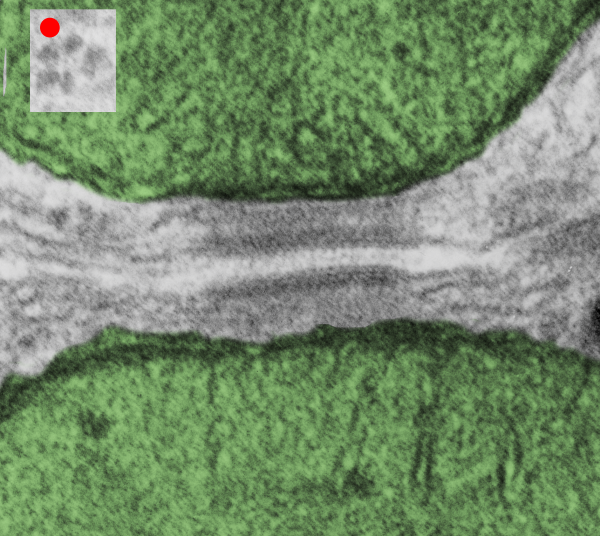 So here is the same electron micrograph as yesterdays post but the mitochondria are actually fuzzed green in a separate layer. Red dot=a ribosome, approximately 27 nm in diameter. The absolutely regular arrangement of cadherins is seen in the middle of the desmosome…. no knots, no tangles, no mess…. just regularly spaced. This micrograph is not retouched to emphasize anything…. it is just the way it looks on the negative. The plasmalemma of the two cells is not in exact cross section and so it is fuzzed. The span between densities both on the plasmalemmal sides and the center densities of this desmosome work out to be about 1/13-14nm. I have not put that together with any models yet and in the image below just sized the molecular structure according to the TEM, not using the actual size (just using the shape).
So here is the same electron micrograph as yesterdays post but the mitochondria are actually fuzzed green in a separate layer. Red dot=a ribosome, approximately 27 nm in diameter. The absolutely regular arrangement of cadherins is seen in the middle of the desmosome…. no knots, no tangles, no mess…. just regularly spaced. This micrograph is not retouched to emphasize anything…. it is just the way it looks on the negative. The plasmalemma of the two cells is not in exact cross section and so it is fuzzed. The span between densities both on the plasmalemmal sides and the center densities of this desmosome work out to be about 1/13-14nm. I have not put that together with any models yet and in the image below just sized the molecular structure according to the TEM, not using the actual size (just using the shape).
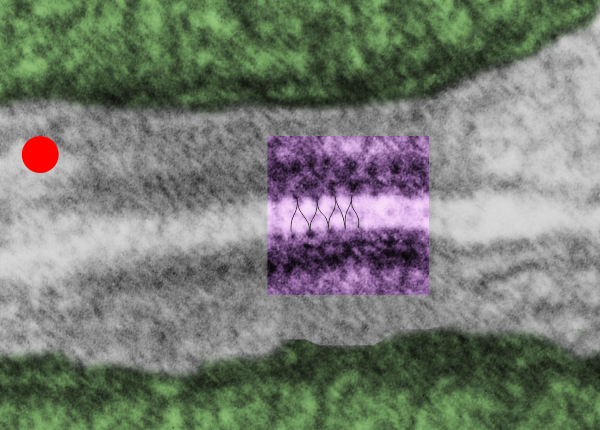

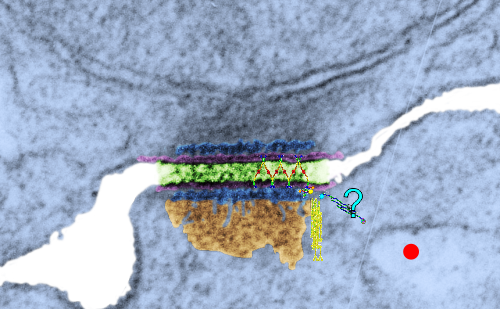
So in the second figure you can see an area that i enlarged, cut and pasted in photoshop, and enhanced in contrast and colored purple and ADDED what i think are the lines of the wishbone arrangements of the cadherins. Densities (increased in contrast here) work out to about 1 for every 13-14nm.
Into two enlargements (one the purple box above and another with the molecular model of a cadherin dimer, copied and mirrorred, fit really nicely into what might be a flexible, stretchy springy type portion of the desmosome. Just a thought…. ??
6110_5080_mouse_f_liver_36,200x_4x
Cadherin molecules overlying a desomosome – mitochondrial tethering
I don’t know much about desmosomes, but they do make unique junctions with nuclear pore intermediate filaments and also very definite connections with mitochondria. The mitochondria – intermediate filament connections with desmosomes show a couple nice ultrastructural changes from the routine: 1) the mitochondrial outer membrane is flattened, or straightened in the area of connection with the intermediate filaments over the desmosome, and it is also a little darker. and 2) the mitochondrial body itself is drawn towards (at least that is the way it looks… as if there was a dragging force) the desmosomal intermediate filaments. The central line of the desmosome is not always visible when i see these tethers (often with two mitochondria in adjacent cells and a single desmosome, but there are molecular models of the cadherins (several) that I cut pasted masked and reduced to fit in the appropriate intercellular space where they would appear. It is a good lesson is relative “size” and “shape”.
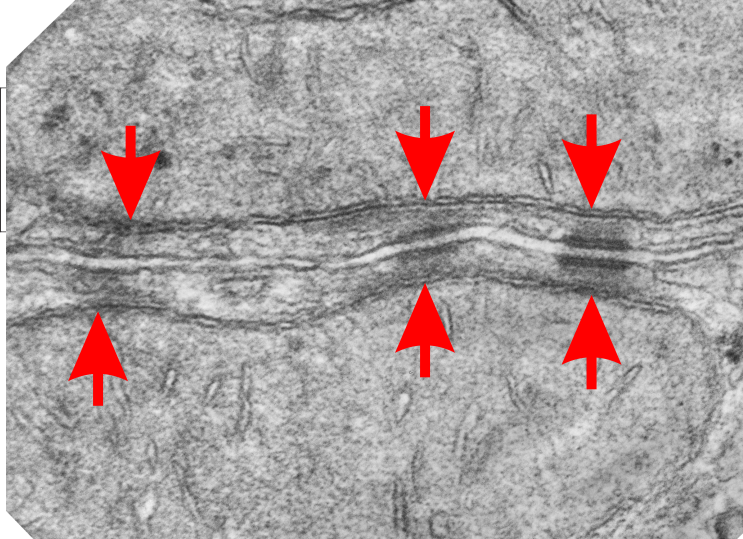
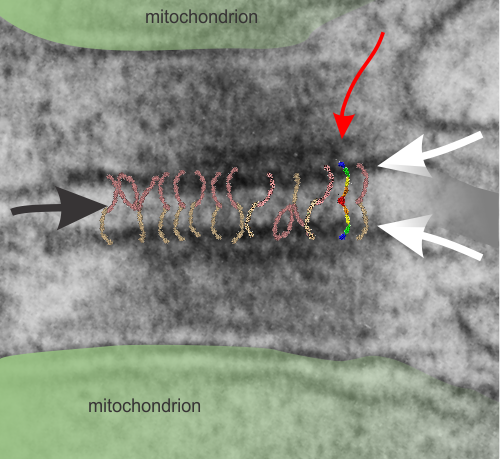
Micrograph: Pale green semitransparent mask is over two mitochondria, one each in two adjacent cells. Red arrow points to the inner dense layer (maybe i will add the known proteins there “in scale), the white arrows point to the plasmalemma of the two separate cells, the black arrow points to the center dense “knotted” region, (maybe i will measure distances between visible knots) and the molecules (several taken from the internet, look similar and were pasted into the intercellular region (see demarcation from right hand side transparent grey.
Triple desmosomal mitochondrial tethers
Both sides of two adjacent hepatocytes show two mitochondria each linking to three desmosomes. Nice to see such symmetry and the implications are quite amazing, in that whatever is happening at the plasmalemmae of these cells is happening “together”, really nice intracellular communication. Red arrows point to the inner mitochondrial membrane, just above where the intermediate filaments of the desmosomes approach the outer mitochondrial membrane. I wish i could say that I saw some cross sections of intermediate filaments here, but i didn’t. The double desmosomal mitochondrial junction on the left is more tangential than the one on the right. The pair in the middle is intermediate.
Intermediate filaments tethering desmosomes and mitochondria
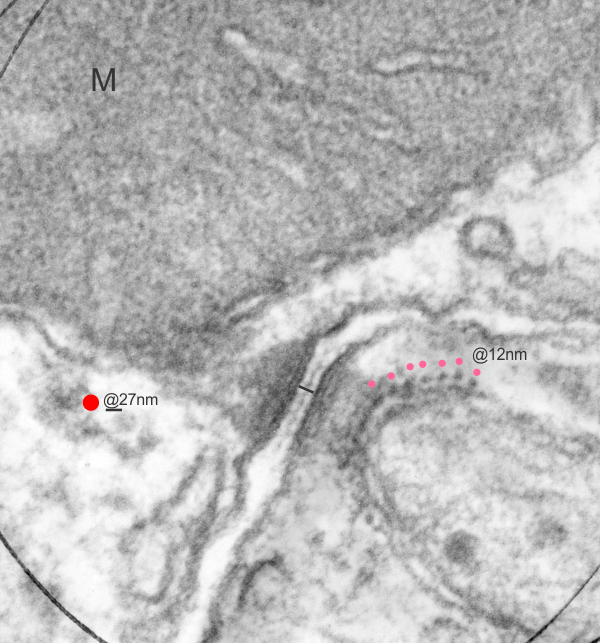
I have mentioned this phenomenon before but i keep finding images which speak to the prevalence of mitochondria (in this image two mitochondria are tethered to a single desmosome, one each in two adjacent cells). in addition to the fun double tether here, look at the “spots” which round the corner of the smaller mitochondrion…. tiny dots of what I measured as about 12nm, but the intermediate filament on cross section is 8-10nm, therefore my guess is that these wre cross sections of intermediate filaments. I did enhance these dots in any way except to change the contrast of the lower half of the image using photoshop. The dots ” cross section of evenly spaced intermediate filaments” are very prominent on their own. Pink dots are proximate to the filament x-sections, the red dot is an approximate ribosomal diameter. 30nm bar in middle is about what the desmosome measures, oriented perpendicular to the two plasmalemmae. N 6105 Block 5080 Mouse, female, orig mag 36000