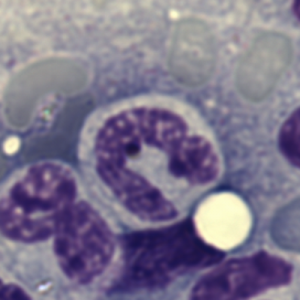
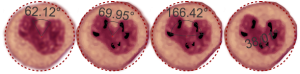
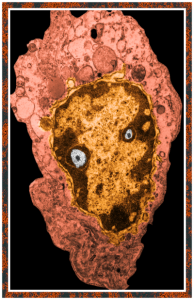
Totally unnecessary post of bone marrow nuclei: but what it says very clearly is that symmetry prevails, may not be the symmetry we expect from a set of identical chromosomes all lined up in on-off status, both sides the same (X chromosome or barr body excepted). I see here a great symmetry, right to left mirror, top to bottom asymmetry, pointy at the bottom because this is an area where the nuclear membrane might just get segmented into a thin band between two bottom areas of chromatin…. going from what might appear to be a point, to two cheeks.
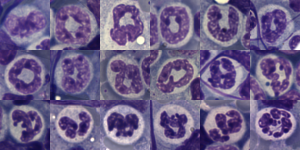
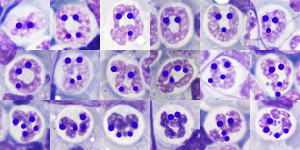
I am thinking about writing up an editorial on this symmetry…. make the cover submission first, always my motto. Each of the hearts below is from bone marrow from an hypoplastic animal (CyP1a1null) either given benzo(a)pyrene or control diet. These particular nuclei are not yet hypersegmented… just fun examples of symmetry, which i have outlined in an identical micrograph below this.
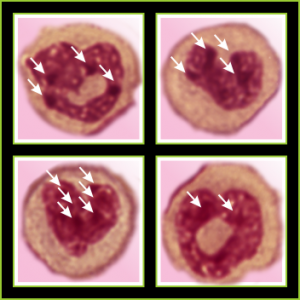
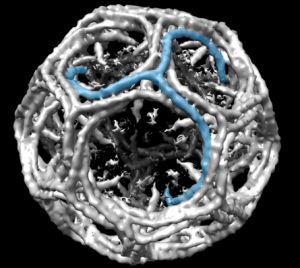
 So there is symmetry, it is bilateral in one dimension, but will take some thinking to see how to describe a third dimension in symmetry.
So there is symmetry, it is bilateral in one dimension, but will take some thinking to see how to describe a third dimension in symmetry.
Because the chromosome territories likely (per published data) kind of interdigitate, most assuredly, at many sites along each chromosome for functional reasons) the symmetry will be highly simplified in light micrographs at best.
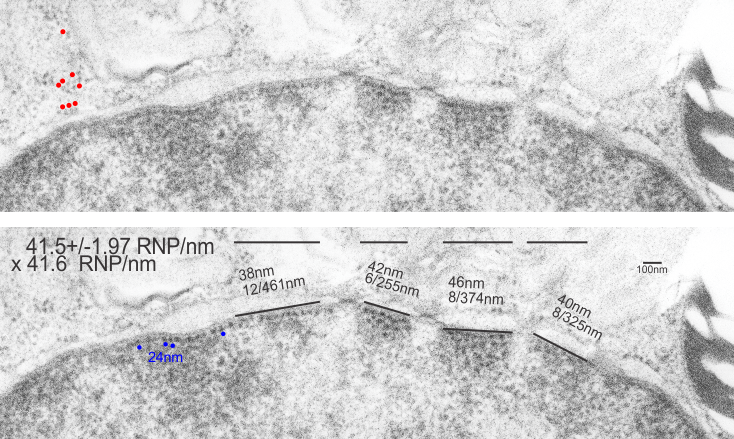
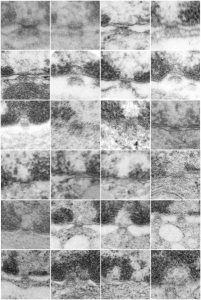
All this discussion could lead to a place to examine the chromatin patterns in the X (barr body) in neutrophils where they are hanging on by a nuclear membrane thread – so to speak) and other parts of the neutrophil nucleus where transcriptional activites are going on.
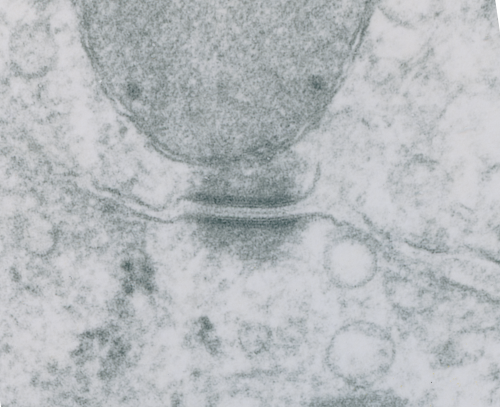
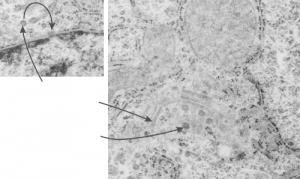
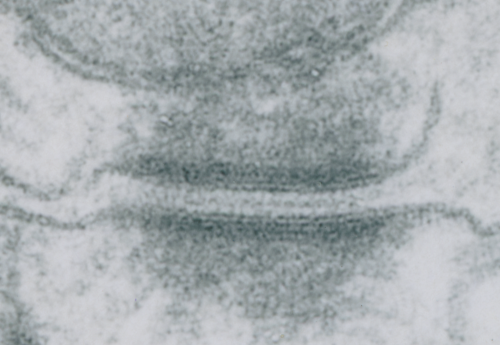 I looks in this image like the plasmalemma of each cell has a very rigid separation of the two leafelets.
I looks in this image like the plasmalemma of each cell has a very rigid separation of the two leafelets.