Human mitochondrial acetoacetyl-CoA thiolase: (found on the RCSB PDB – 2F2S)
 This site is awesome, and I cant even begin to imagine how many smart people have worked tirelessly to make these data available to the average joe like myself. Thanks to them.
This site is awesome, and I cant even begin to imagine how many smart people have worked tirelessly to make these data available to the average joe like myself. Thanks to them.
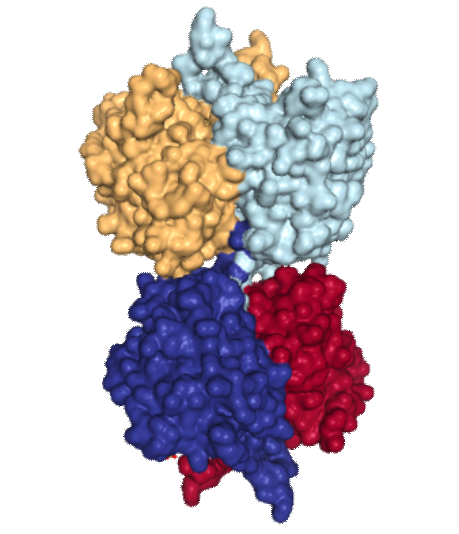
The image of acetoacetyl-CoA thiolase of four different nearly identical chains identified by color in image to right. I did not find articles (except one) stating that this was a membrane protein, and in which (outer or inner or cristae membranes) it would reside, and whether at the cristae pore or not. My interest in posting pictures of this protein came after seeing that it has a 3dimensional symmetry that was visually quite wonderful. Below are two sets of images taken at points of symmetry using the RCSB PDB link listed above. The symmetry is best seen with the ribbon molecules, but also is obvious with the space-filling and residue coloring modes. I found this protein while looking for other mitochondrial membrane proteins, I am struck with how often nature mirrors flips and rotates basic protein elements to create function. In the images below molecules are colored as N-terminus (blue) to C-terminus (red); from 5′ end (blue) to 3′ end (red) — , bottom row is colored by residue. 3 green lines show symmetry (the line in z axis comes straight forward and looks like a point at the intersection of x and y. The green box shows a close vertical mirroring of the molecule…. rotated 90 degrees. I think the particularly interesting loops (four total) would have some special functions, seen at the top and bottom as presented in the ribbon diagrams.


Last image shows hydrophobic sites (right)… doesnt look like there is a region that would be membrane-bound does it?
It might sound silly to try to find out the size and shape of the known membrane proteins in mitochonria but just looking at the TEM images of mitochondria that I have accumulated over many years and many experimental conditions i am convinced that I can find some unique things to describe and show, but not knowing enough about the membrane proteins makes it less than satisfying. So at the beginning of the search, the ATP synthases was clearly fun, and easy to spot in some abnormal mitochondria.








