I have scoured the literature to find any images, and even diagrams, of rat native and recombinant surfactant protein D, hereafter called SP-D. There are some very nice images (perhaps the best) by Crouch et al, published in JBC, (also using recombinant SP-D) which I have cut and pasted into this video, counting all the trimeric arms with “attached” carbohydrate recognition domains. Crouch et al indicated that about 5% of the structures are multi-assemblies, that is something around 12 arms? or more is what they show. Actually two authors have commented on the fuzzyball rarity in shadowed images for microscopy. Other images came from google searches for SP-D diagrams, and many of these are from McCormack, and from others in the lab here at Cincinnati’s Childrens Hospital.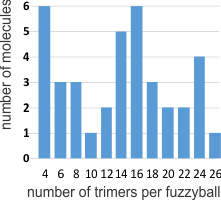
The three dimensional structure (multimers) of SP-A and SP-D have been shown to be important for function. Vieira et al 2017“In conjunction, these data demonstrate the critical role of oligomeric and tridimensional structure of surfactant protein A and surfactant protein D for proper function.” and also say “SP-A and D can target pathogens by simple aggregation, without a direct interaction with the cell. Similar to complement C1q, SP- A can function as an “activation-ligand” which facilitates particle uptake once it is coated by Immunoglobulin G”
SP-A and SP-D have distinct functions though both are quite easily given the “title” of multivalent innate immune proteins. What is interesting is that under some circumstances and not others, they may adopt a more spherical and highly oligomerized state, or they may remain trimers, hexadecamers, dodecamers and in lower oligomerization states. For some reason the bouquet for SP-A as an 18-mer is most frequently cited in the literature when in fact the electron microscopy might show full fuzzyballs of SP-A, and other forms (periodicity in flat sheets, under other circumstances. In the same micrographs of type II alveolar cells, SP-D fuzzyballs really havn’t been encountered, but in vitro SP-D makes great fuzzyballs.
https://thankuscience.com/wp-content/uploads/2018/09/SP-D.avi (cut and paste) SP-D .avi link
I was curious whether there was a pattern to the number of trimers in any given fuzzyball and for SP-D there were numerous articles which actually had pictures of oligomers. The number of arms (trimers) for each fuzzyball of SP-D, randomly encountered in publications (not to be confused with randomly photographed under the microscope… as i am quite sure there was selection bias in photographing these) is shown on x axis of this quick summary graph. Incidence is on the Y axis. I could have counted more (and may do that) in some images where there was indication that perhaps there were arms and CRDs that had been displaced during collapse of the fuzzyball onto the grid. Image below is taken from Crouch et al mentioned above and used without permission.
Here is a short videoclip of SP-D fuzzyballs — you can see where I have counted the carbohydrate recognition domains, as I overlayed them on the existing micrograph with red dots. There are places where I might add or subtract one now and then from what is pictured, but in general you can see that top number is something around 26 arms (each a trimeric molecule) in some fuzzyballs. I think more than 32 (a multiple of 4 is likely) arms in any given fuzzyball is going to be pretty uncommon. It is very apparent in some that groups of 4 arms persist. While the orientation of SP-D arms is probably end to end in those groups of 4, I am thinking that SP-A fuzzyballs are going to be four groups of 18-mers. Whether such is found in the literature I will soon find out.