It is nice to read that something is measurable, that has a dimension in nm that one can put into some kind of life-perspective. I don’t know how accurate those individuals were in giving a distance measurement to their images are, and I am giving each the benefit of the doubt in terms of attempting to be accurate) i guess they are quite accurate. I know for myself, using three or four different electron microscopes, and various different paper products, films, dryers, etc, that there is shrinkage, stretching, and all sorts of changes that have occurred over time (even 35 years) that affected the dimensions of objects on the micrographs, even though by a very small amount.
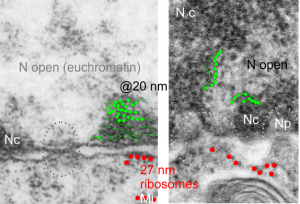
I am using 27nm as an internal control for magnification+enlargement of images. In this case, the 27nm red dot, in all images has been derived from about a dozen ribosomes, and labeled as such. The green dots are about 18-20nm in diameter in these micrographs, and strung as if “beads on a string” (not the beads on a string of nucleosomal DNA, but production of preRNA in the nucleus?) but is it not strange that there is a repeated motif – that is- beads on a string- showing up a lot, both in the nucleoplasm, the nucleolus and the cytoplasm (ribosomes on mRNA) as a kind of “convenient” little pattern. And who knows where else this pattern might be found). So the green dots show this motif, in a parallel-linear pattern, in the condensed (or closed compartment) chromatin. Both of these examples come from Gclc ko mouse hepatocytes. Red dots are ribosomes.