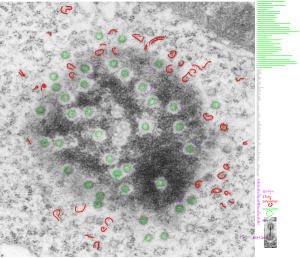
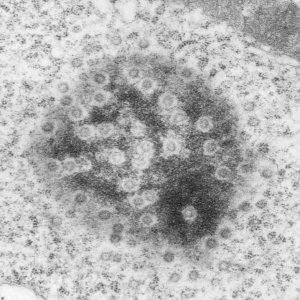
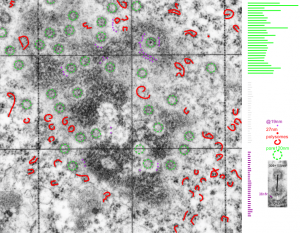
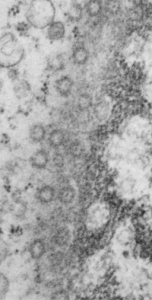
16040 65718 liver 14Cos 24 hr no NTBC, unidentified cell, not likely an hepatocyte, maybe an interstitial cell. Mitochondria in adjacent cells top left and right have condensed mitochondria. One of the interesting things about the “inclusion” in this nucleus is that it does not resemble any cytoplasmic structure in the cytoplasm of that cell, in fact the cytoplasm of this cell is quite different with lots of vacuoles (lower left) and an ill-defined shape, on one side very scant cytoplasm, which makes it less likely to be an hepatocyte than an interstitial type cell.
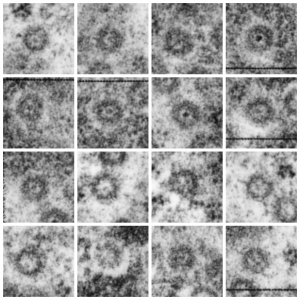
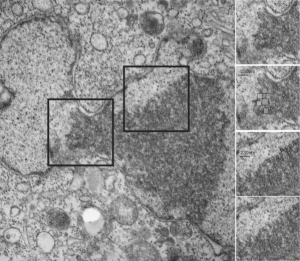
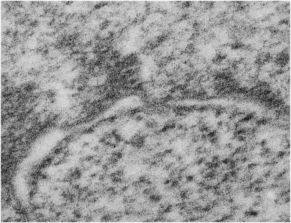
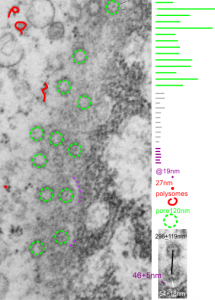
Three areas along the margin of the inclusion have been cut and enlarged (denoted by the three boxes in the original micrograph). The inset at the top shows there is really no distinct double membrane typical of the nucleus (inner and outer membranes and nuclear pores) and this might be a tangential cut-artifact. In the inset from the middle left, theer is a band of chromatin along the left border of the inset but again, no double membrane typical of nucleus. The inset from the lower right abuts up against nucleuplas which is very electron lucent, but along this area there is a density to the inclusion which looks like it might be some kind of lamin proteins.
Another great feature is the constriction appearing in lower middle of the inclusion, densities that look like they might have been some kind of “ring” around part of this structure.
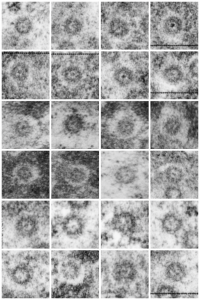
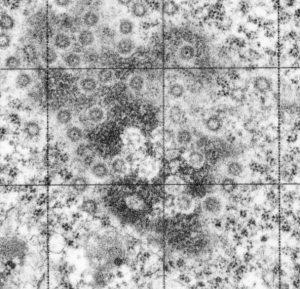
Ribosomes (estimated at 27 nm in diameter from the cytoplasm have been enlarged with the insets (also shown in the original). Bar is 10x the ribosomal diameter at approximately 270nm.
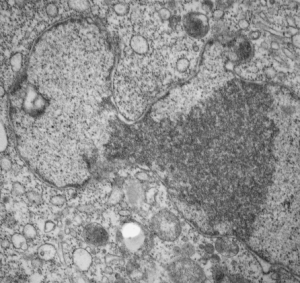
The inclusion/invagination is full of robosomes and RER membranes (I guess that is what they would be named) with an homogeneous appearance and the vesicles or RER appear to be dilated to about the same uniform degree.
There is a nucleolus just above the inclusion. Animals with genotype undergo a pan-apoptosis among hepatocytes within a few days of birth, this doesn’t look like apoptosis yet, and also doesn’t appear to be an hepatocyte.



 .
.