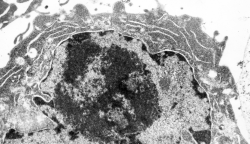
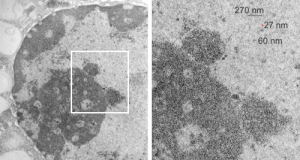
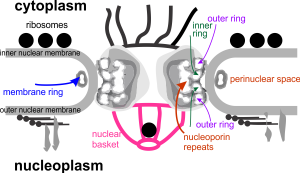
Looking at the sum total of nuclear architecture seen from the viewport of the electron microscope, I am trying to summarize the topology and/or segregation of protein-DNA, protein-RNA and protein-protein domains therein. It seems that many names have been associated with very many small structural entities within the nucleus, (and nucleolus), and not all these molecular studies have been diligent about identifying them with transmission electron microscope (actually using fluorescent probes and light microscopy gives pretty pictures, merging reds and greens — but this is still like looking at the barn door when one wants to see the insects on the wood) thus making it a little difficult to interpret their results. I did find an early manuscript, from 1963, which helps, published just about 10 years after the commercial availability of the electron microscope (yes I laughed since the author of the manuscript named the equipment, an old Seimens 1 — the microscope I used for 30 years was the next generation, the Seimens 1A. yep, old).
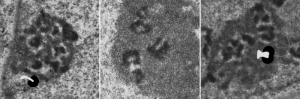
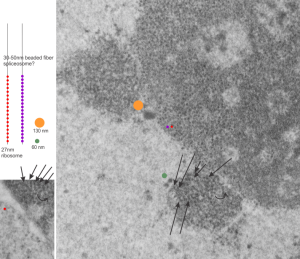
The publication is by Nicole Granboulan and appears to be readily available for perusal and it focuses on the nucleolus, a portion of the nucleus dedicated to ribonucleoprotein biogenesis. I have taken portions of three micrographs, an uninfected, and two SV40 infected, and put them side by side to show the changes in the nucleolus as it becomes a production site for viral pre-ribosomes as shown by the great increase in the granular area as time after SV40 infection increased. There is an augmentation to the dense fibrillar compartment, which initially to me looks like a crescent moon with the fibrillar centers poking in like mushroom stalks, to feed the production of viral RNA. It is clear that the surface areas between the fibrilar centers (where the rDNA genes are located) and the inside of the crescent moon portion of the dense fibrillar areas has occurred as well. An increase in the granular component in uninfected nuclei (uninfected cells) is also occurring when cells are in S phase of the cell cycle.
The black and white diagram overlays with white arrows show the change from U to M shape of the dense fibrillar regions, and the white areas of the cartoon, being the fibrillar centers.