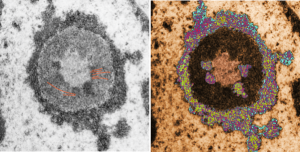
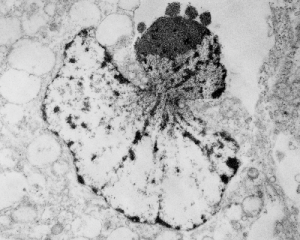
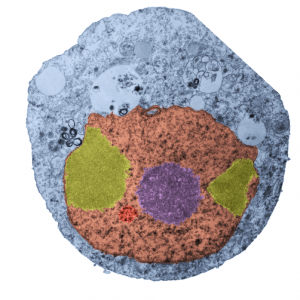
While interchromatin granule clusters were thought to be the “the same structure” as nuclear speckles, i wonder if that opinion was justified at the time, I am not about to make that leap. I need to sort this out, since with plastic sections stained with toluidine blue, the interchromatin granules really were not “lucent or unstained” areas, but a light tan color, if i remember right, and with TEM they were actually more closely punctate than usual euchromatic nucleoplasm. So the connection needs to be further varified…and this might be why there are mixed notations and confusion and disparate explanations as to what the speckles and interchromatin granule morphologies identify. Also, what I have seen referred to as interchromatin granule clusters have two (at least) maybe three or four sizes of granular elements depending upon metabolic or functional state of the nucleus is (as in G,S and also stages of apoptosis, maybe necrosis as well. (Of course all the (11 so far) listed processes of cell death and cellular-self destruction could be listed here but I just subscribe to the main ones, necrosis and apoptosis for now.) This cell has nucleolus dark blue, nucleus golden, cell cytoplasm greyscale, interchromatin granule cluster of pink pink, and white box shows area enlarged in picture below. The larger (one quite large) and four or so smaller clusters can certainly point to some metabolic phenomenon which is not really occurring in a non-cultured-non-exposed cell.
While interchromatin granule clusters were thought to be the “the same structure” as nuclear speckles, i wonder if that opinion was justified at the time, I am not about to make that leap. I need to sort this out, since with plastic sections stained with toluidine blue, the interchromatin granules really were not “lucent or unstained” areas, but a light tan color, if i remember right, and with TEM they were actually more closely punctate than usual euchromatic nucleoplasm. So the connection needs to be further varified…and this might be why there are mixed notations and confusion and disparate explanations as to what the speckles and interchromatin granule morphologies identify. Also, what I have seen referred to as interchromatin granule clusters have two (at least) maybe three or four sizes of granular elements depending upon metabolic or functional state of the nucleus is (as in G,S and also stages of apoptosis, maybe necrosis as well. (Of course all the (11 so far) listed processes of cell death and cellular-self destruction could be listed here but I just subscribe to the main ones, necrosis and apoptosis for now.) This cell has nucleolus dark blue, nucleus golden, cell cytoplasm greyscale, interchromatin granule cluster of pink pink, and white box shows area enlarged in picture below. The larger (one quite large) and four or so smaller clusters can certainly point to some metabolic phenomenon which is not really occurring in a non-cultured-non-exposed cell.
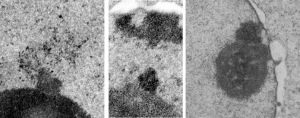
The purpose here is to comment on the various sizes and shapes of such densities within the interchromatin granule clusters, and to examine transmission electron micrographs of so many cell-death projects to see whether concistent patterns interchromatin granule clusters have components change in size, position, density, and shape depending on which processes are occurring within the nucleus.
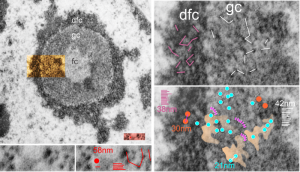
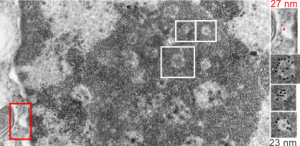
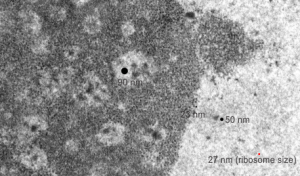
In particular, when A549 cells were examined after knockdown of an antiapoptotic gene (using si67), then the interchromatin granule clusters contained large aggregated densities (large in comparison to the smaller entities in untreated cells). Here I have localized these in a single cell (transmission electron micrograph of the area bounded by the white box above), just to point out what is observed. Relevance is yet to be determined for these larger than usual areas of of conspicuous density within interchromatin granule clusters. The curved arrow points to bar-like organization of a filament or fibrilar area (sometimes this ribbon type linear organization is seen in cajal bodies) and the width of these bar-like fibrilar structures would be something around 30-40 nm, if compared to the size of a ribosome (red dot in the center of the text that says 500 nm). The large structure (electron dense) beneath the interchromatin granule is the top part of the nucleolus seen in the image above. neg 18272 block 78932 A549 cell si67 knock down of antiapoptotic gene in vitro.  )
)