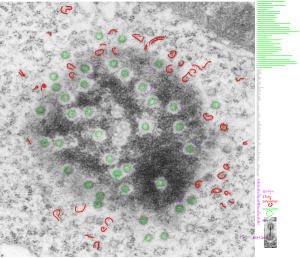
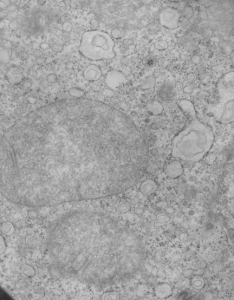
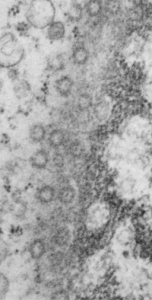
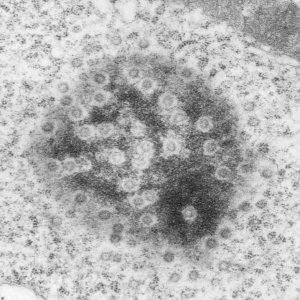
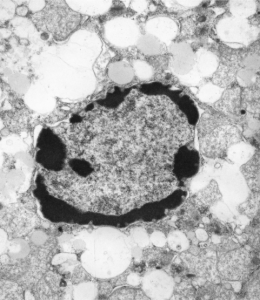
Here are some nice round structures and were they always near the nuclear membrane i would count them as tangentially sectioned nuclear pores since they are sized so similarly (though i have not seen any central densities like i see in nuclear pores where actual proteins are being imported or exported. There are also some profiles of membrane (with contents) that are pretty linear for golgi seen as well, and don’t quite fit that golgi iconic morphology. This is a fetal hepatocyte. Arrows point to pore and to similarly sized rounded structures that are not likely to be tangentially section pores from the nucleus shown.
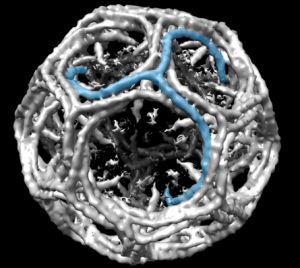
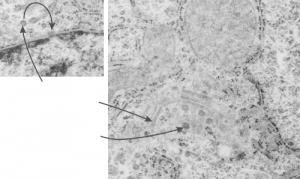 Might be a section of a clathrin coated vesicle…. when i google it, this wonderful model came up. Sure looks (at 100nm) like it could work. What a marvelous amount of research has provided the molecular structure (triskelion) shown by them (on wikipedia) in blue.
Might be a section of a clathrin coated vesicle…. when i google it, this wonderful model came up. Sure looks (at 100nm) like it could work. What a marvelous amount of research has provided the molecular structure (triskelion) shown by them (on wikipedia) in blue.
Category Archives: Electron micrographs of liver
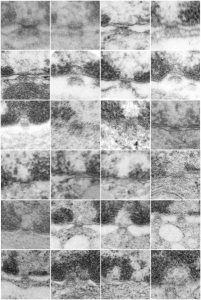
More nuclear pores, just randomly collected various cell types
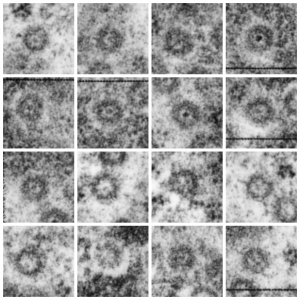
More nuclear pores, just randomly collected various cell types. The criterion that I used was simply the presence of vertical filaments (either part of the nuclear pore basket on the nuclear side or the filaments in the cytoplasm on that side. These come from lung alveolar type II cells, from hepatocytes, and from CoS14 ko mice, rescued and non-rescued, and probably a couple from Gclc conditional KO mice. These are just to give the limit of what effects random sectioning through a block of tissue can do to a round or discoid object. These pores are cut perpendicular (side views), and chromatin is always on top, cytoplasm is always below.
mitochondria and SER: liver
This micrograph has some interesting features for an ostensibly normal mouse liver: normal in that it is part of a study on Gclc loss, which causes early death (specifics and article here) but this liver should not have had so many vesicles within vesicles as seen in the Gclc ko mice, and typically they are vesicles within vesicles without ribosomes. here there are vesicles within the vesicles of RER …17941_73225_liver_Albwc_Gclcii_1mo_no_NAC
Nuclear pore measurements: pore to pore, chromatin exclusion zone, granule to granule
Three measurements taken in a series. Pore to pore, pore to to edge of chromatin (which i found is called the chromatin exclusion zone, so am sticking with that since it clearly is an active event, and may vary with cell type and cell activity) and inter-@19 nm “beads” that are seen as the organized chromatin just adjacent to the nuclear pore (the first thing seen after the chromatin exclusion zone). This micrograph is from the same images as a previous post, which is 16027_65718_14Cos_ko 24hr no NTBC measurements.-2a.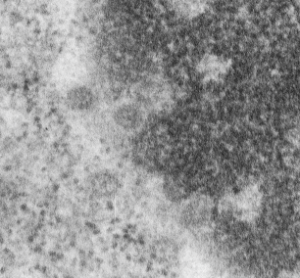
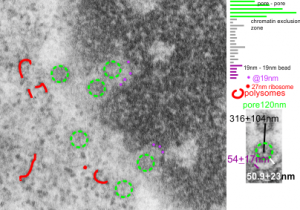 n=5, pore to pore=316nm+104nm; n=15, chromatin exclusion zone, 50.9+23nm; n=5, 19nm to 10nm chromatin granule, 54nm+17.7nm
n=5, pore to pore=316nm+104nm; n=15, chromatin exclusion zone, 50.9+23nm; n=5, 19nm to 10nm chromatin granule, 54nm+17.7nm
More nuclear pore measurements
This hepatocyte was from a mouse (#5) which was a 14CoS nujll that did not receive NTBC as a rescue drug. This is 24 hours into the neonatal period. Nuclear pores have fewer central densities (presumably being proteins transported). I am measuring for distance beside the nuclear pore that the chromatin exclusion zone hoping that differences will show up with various conditions in previous experimental models.
Top micrograph, unretouched, bottom micrograph, pores used in calculation have circles, red dot = a 27nm ribosome, purple dots are areas of the chromatin just adjacent to the chromatin exclusion zone surrounding the nuclear pore.
Nuclear pores and polysomes and other things
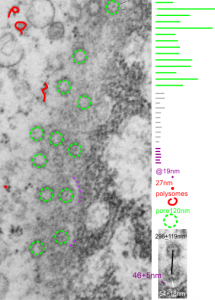
Continuing an evaluation of nuclear pores in various experiments from the past, here is a microgaph from 17709_65053_14CoS-/- NTBC 1yr. I used some of the opportunely sectioned “almost top down” cuts of nuclear pores to look at the inter-pore distance (which here measures , the chromatin exclusion zone, and the distance between the adjacent chromatin areas that have that “beads on a string” look. Distances are derived from an approximate size of the ribosome at 27nm in diameter, nuclear pore diameter of 120nm, and a 19nm diameter for the adjacent chromatin at the edge of the chromatin exclusion zone.
Unretouched photograph of portion of a nucleus from an hepatocyte from CoS-/- mouse maintained on NTBC for 1 year. and the marked-up micrograph below.
The nuclear exclusion zone is 54nm+18, the inter-pore distance is 289+119nm, and the distance between the densities in the chromatin (purple dots) of the edge of the chromatin exclusion zone is 46+5. Red curly-cues are polysomes, red dot is one ribosome (taken at 27 nm). Green 8-edged rings are nuclear pores. (bottom picture)
Measurements of areas surrounding the nuclear pore: hepatocyte
Some measurements (see previous post for comparison data), might be useful in a relative sense to others looking at nucleus, nuclear architecture, and nucleolar architecture.
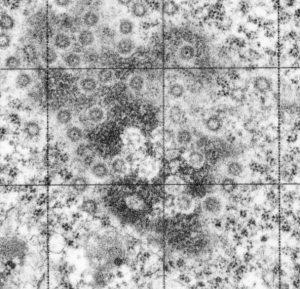
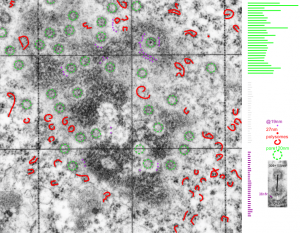
This particular image is unretouched tangential section of an hepatocyte nucleus (17529neg73229block, anm203,wcii -Gclc 28d). There are numerous nuclear pores sectioned tangentially in this image, about 35 of which are there 15 or so in sufficient detail to demonstrate a density within the center of the nuclear pore demarcating the site that pre-mRNA and larger particles are ushered through (both ways) (about 44%). Compare this micrograph with a previous post of a similar tangential section for parallel measurement. They are not identical, but close. Curley red lines demarcate polysomes.
In this micrograph the pore to pore distance is n=34 x=482+177nm, the distance from the outer part of the nuclear pore to adjacent chromatin is n=97, 62.7nm +/-32nm, and the distance between the 19-20 nm beads on a string in the chromatin next to the chromatin exclusion area is n=39, 46nm +/- 13nm. The actual measurements are shown at right (green=pore center to pore center), grey=chromatin exclusion areas, that is the distance between the edge of the nuclear pore and the adjacent chromatin, grey, and purple, the distance between the 20nm beaded-periodicity found in the pore-adjacent chromatin.
Nuclear pore gallery-1
A collection of nuclear pores (top down) from a single tangential section of an hepatocyte nucleus from a mouse +/- for 14CoS (as opposed to the null). These show that about three quarters of the nuclear pores have some kind of transport item (like a pre-ribosome, or something else) being imported or exported all the time. Just afun look at these adjacent pores. (excuse the morphometry grid from my inkjet printer back in the days when point counting was the method of choice for gathering numerical data from TEM images.
So these from a single cell show something about morphology… and the distance of the heterochromatin exclusion area surrounding the pore and the inter-pore distance and the size of the chromatin “beads on a string” in the adjacent chromatin I posted before.
Surrounding elements to nuclear pores: measurements
Some measurements, just made for my own information, might be useful in a relative sense to others looking at nucleus, nuclear architecture, and nucleolar architecture. Some data exist of course, but In the dataset that I am putting together from many different experimental protocols (archived) might lead to some new insights on numerical density of nuclear pores during times of cell stress, apoptosis, and in previously studies KO mice. The numbers are meant to be “observations” and a suggestion.
This particular diagram has as its background (top image) unretouched, and from a group of micrographs which were used for morphometry (hence the ink jet grid on the surface). It is hepatocyte, from a +/- 14CoS mouse which did receive NTBC (as a control for experimental mice receiving NTBC (data here) but showed pretty normal histology – but I predict with enough time and scrutiny something would show up. Octameric nuclear pores are shown as green dotted circles, ribosomes and polysomes presumably attached to the outer nuclear membrane, red, ribosomes, red = @27 nm diameter, peri-pore chromatin beads diameter, @19-20nm. Condensed chromatin at the inner nuclear membrane is well defined and “wound” in many places as a fibril also, possibly around 20+ nm diameter but longer lengths. About %66 of nuclear pores in this photo have central densities indicating some molecule is in transport one way or the other.
This particular animal had a great tangential section showing a large portion of the nucleus studded with nuclear pores and I used that opportunity to make some measurements on the diameter of the chromatin with beads (@19nm)-on-a-string appearance adjacent to the nuclear pores but outside the chromatin exclusion area around the pores and the distance between those beads (38nm); to measure the width of the chromatin exclusion area at places around the nuclear pores (maybe 2 -5 measurements perpendicular between pore and chromatin, per pore) (60nm+ 7nm and to measure the pore to pore distances as well (n=27, MEAN, 309nm+ 109nm (SD)).
Endstage apoptosis in hepatocytes
Here is a textureless (or nearly so) grouping of DNA around the periphery of an hepatocyte from a 14CoS ko hepatocyte, from a young mouse not rescued with NTBC. Symmetry to clumps or peripheral digested DNA is obvious. There remains some substructure to the “euchromatin” in the nucleus as well as some structure to the areas of presumptive digested chromatin.