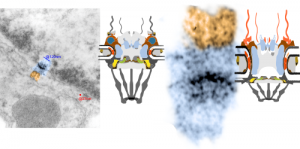
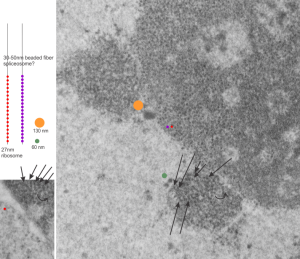
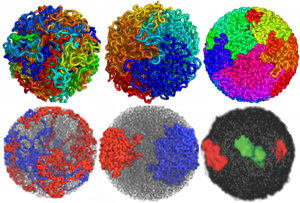
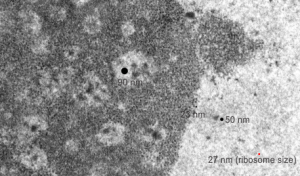
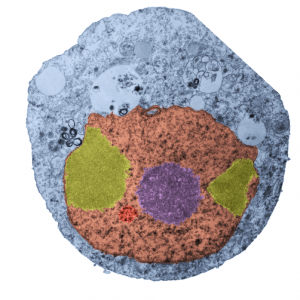
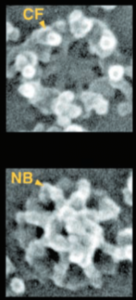
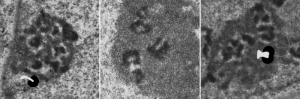
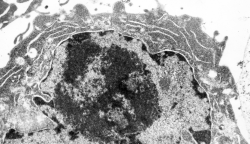
I ask forbearance for my posting this. I have seen this asymmetrical type of apoptosis in some experiments that were done while investigating an anti-apoptotic gene. I was just doing the electron microscopy, others were doing the cell culture and knockdown experiments. Typically apoptosis in culture presents with a round nucleus lots of large nucleoli and a bilateral (mostly) distribution of granular material along the inner nuclear membrane of the cell as it dismantles itself. The squeezing and pinching in this type of apoptotic cell was just really unusual and as is always the case on government grants, the reasons were never flushed out because of time and money constraints.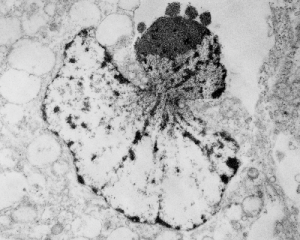 I bet there is a really interesting reason for these, and this particular set of culture dishes got a scrambled version of an si67 probe and was supposed to be a control. It seems to me that part of the nucleus a just got extruded? I did count 8 nuclear pores (main criterion being an obvious set of cytoplasmic filaments off the cytoplasmic side of the pore. Few mitochondria, many vesicles. One interesting thing about this apoptotic style is that the cytoplasm is not full of ribosomes as occurs in apoptosis in many cell types.
I bet there is a really interesting reason for these, and this particular set of culture dishes got a scrambled version of an si67 probe and was supposed to be a control. It seems to me that part of the nucleus a just got extruded? I did count 8 nuclear pores (main criterion being an obvious set of cytoplasmic filaments off the cytoplasmic side of the pore. Few mitochondria, many vesicles. One interesting thing about this apoptotic style is that the cytoplasm is not full of ribosomes as occurs in apoptosis in many cell types.
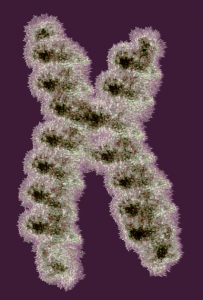
It is not something i could have made up (LOL) but my initial guess is that the four little ball-antennae are some remains of cajal bodies, and heretically i could suggest that the linear divisions radiating out into the “skirt” of whats left of the nucleus, likely has something important to do with each chromosomal territory (separations, walls, scaffolding, call it what you like).