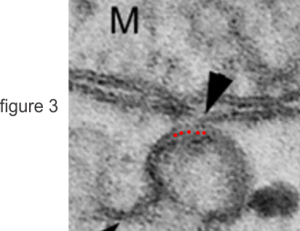
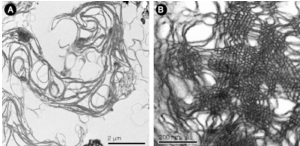
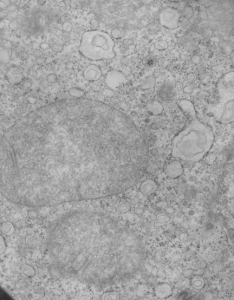
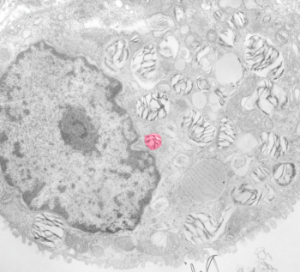
These nice interactions between nuclear pore outer filaments and the outer membrane of the mitochondrion have intrigued me for at least 3 decades. They first came to my attention when looking at liver, hepatocytes specifically, where they are frequent, and uncluttered by a lot of other cell machinery and organelles. It seems pretty obvious that their tethering an energy source to fuel translocation of cytoplasmic proteins into the nucleus: “proteins (such as DNA polymerase and lamins), carbohydrates, signaling molecules and lipids moving into the nucleus” according to wikipedia. The author of the wikipedia post suggests that 1000 translocations per pore complex occur each second. Smaller molecules diffuse through at the peripheral portions of the pore and specific nuclear localization sequences are recognized by the nucleoporin proteins to assist larger molecules through the central part of the pore channel. That author also suggests that the pore transport does not require energy, so there must be some function for the mitochondria found associated with nuclear pore complexes (and sometimes appear to be just bent and dragged toward them). It is possible that assembly and disassembly of nuclear pores would require mitochondria. It has been suggested that three NUPs which do occur on the outer (cytoplasmic) filaments get phosphorylated. It is likely a testable hypothesis, which could be ferreted out at least a little in examining different cell types and relating that to mitotic index, or alternatively, from unperturbed to apoptotic state. It is not that unusual to find three different areas of the same mitochondrion to be seen associated with outer nuclear pore filaments.
One mitochondria (on the left) shows two such close encounters. These examples come from either alveolar type II cells or hepatocytes; grey dots 27 nm ribosome, bar=270 nm. It is interesting that the nuclear basket (filaments on the nucleoplasmic side of the nuclear pore complex) are not visible in any of these pore-mitochondrial associations, except maybe slightly in image on lower right. In each micrograph, the mitochondrion is top side, a portion of the nucleus is below.