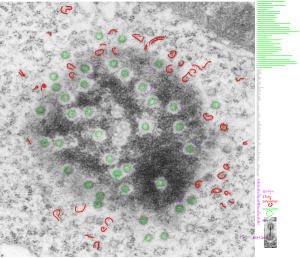
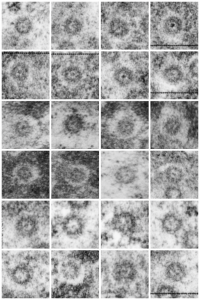
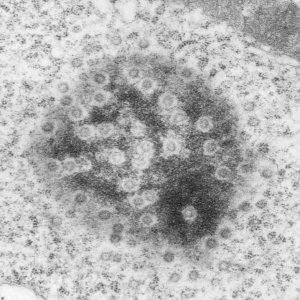
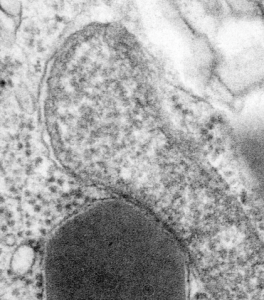
This post is just for fun, as when i was looking at some electron micrographs of the Gclc mouse liver study I found this critter, just with big eyes (one of which i added the pupil to using photoshop — my apologies) but otherwise not retouched. The exact data are neg17894_block74138_animal706_wcii_NAC. Initially I was interested in the nucleoli, but naggingly amazed about the “devil” appearance of the nucleus as a whole.
Color was of course added using photoshop, all else remains the same. When considering what to call this character i googled “names for satan” and behold, the list of about 66 names from various cultures and groups showed up (cut and pasted from Wikipedia (thank you wikipedia) (below).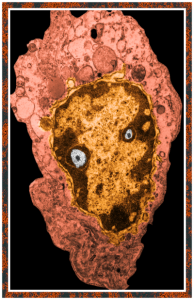
Abaddon—(Hebrew) the destroyer
Adramalech—Samarian devil
Ahpuch—Mayan devil
Ahriman—Mazdean devil
Amon—Egyptian ram-headed god of life and reproduction
Apollyon—Greek synonym for Satan, the arch fiend
Asmodeus—Hebrew devil of sensuality and luxury, originally “creature of judgment”
Astaroth—Phoenician goddess of lasciviousness, equivalent of Babylonian Ishtar
Azazel—(Hebrew) taught man to make weapons of war, introduced cosmetics
Baalberith—Canaanite Lord of the covenant who was later made a devil
Balaam—Hebrew devil of avarice and greed
Baphomet—worshipped by the Templars as symbolic of Satan
Bast—Egyptian goddess of pleasure represented by the cat
Beelzebub—(Hebrew) Lord of the Flies, taken from symbolism of the scarab
Behemoth—Hebrew personification of Satan in the form of an elephant
Beherit—Syriac name for Satan
Bilé—Celtic god of Hell
Chemosh—National god of Moabites, later a devil
Cimeries—Rides a black horse and rules Africa
Coyote—American Indian devil
Dagon—Philistine avenging devil of the sea
Damballa—Haitian Vodou serpent god
Demogorgon—Greek name of the devil, it is said should not be known to mortals
Diabolus—(Greek) “flowing downwards”
Dracula—Romanian name for devil
Emma-O—Japanese ruler of Hell
Euronymous—Greek Prince of Death
Fenriz—son of Loki, depicted as a wolf
Gorgo—dim. of Demogorgon, Greek name of the devil
Haborym—Hebrew synonym for Satan
Hecate—Greek goddess of the underworld and witchcraft
Ishtar—Babylonian goddess of fertility
Kali—(Hindu) daughter of Shiva, high priestess of the Thuggees
Lilith—Hebrew female devil, Adam’s first wife who taught him the ropes
Loki—Teutonic devil
Mammon—Aramaic god of wealth and profit
Mania—Etruscan goddess of Hell
Mantus—Etruscan god of Hell
Marduk—god of the city of Babylon
Mastema—Hebrew synonym for Satan
Melek Taus—Yezidi devil
Mephistopheles—(Greek) he who shuns the light, q. v. Faust (Greek)
Metztli—Aztec goddess of the night
Mictian—Aztec god of death
Midgard—son of Loki, depicted as a serpent
Milcom—Ammonite devil
Moloch—Phoenician and Canaanite devil
Mormo—(Greek) King of the Ghouls, consort of Hecate
Naamah—Hebrew female devil of seduction
Nergal—Babylonian god of Hades
Nihasa—American Indian devil
Nija—Polish god of the underworld
O-Yama—Japanese name for Satan
Pan—Greek god of lust, later relegated to devildom
Pluto—Greek god of the underworld
Proserpine—Greek queen of the underworld
Pwcca—Welsh name for Satan
Rimmon—Syrian devil worshipped at Damascus
Sabazios—Phrygian origin, identified with Dionysos, snake worship
Sammael—(Hebrew) “venom of God”
Samnu—Central Asian devil
Sedit—American Indian devil
Sekhmet—Egyptian goddess of vengeance
Set—Egyptian devil
Shaitan—Arabic name for Satan
Shiva—Hindu the destroyer
Supay—Inca god of the underworld
T’an-mo—Chinese counterpart to the devil, covetousness, desire
Tchort—Russian name for Satan, “black god”
Tezcatlipoca—Aztec god of Hell
Thamuz—Sumerian god who later was relegated to devildom
Thoth—Egyptian god of magic
Tunrida—Scandinavian female devil
Typhon—Greek personification of Satan
Yaotzin—Aztec god of Hell
Yen-lo-Wang—Chinese ruler of Hell
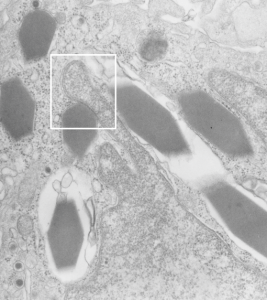
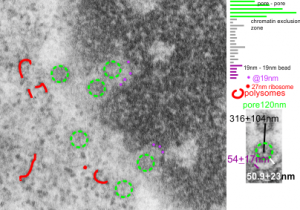
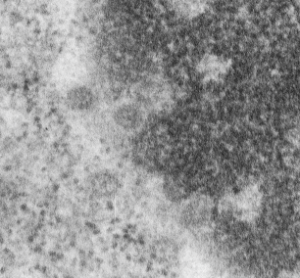
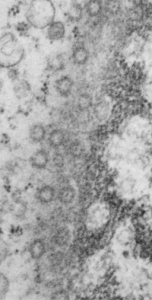
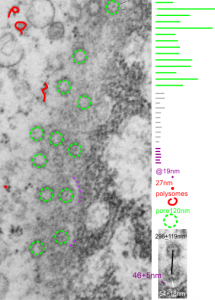
 I can visualize these little A-frame things with heads on them about 10 of them on the inner nuclear membrane. Outer nuclear membrane and granule membrane is really indistinct… owing to the ribosomes in the limiting membrane of the crystal, one could even assume that this crystal might be in the perinuclear space.
I can visualize these little A-frame things with heads on them about 10 of them on the inner nuclear membrane. Outer nuclear membrane and granule membrane is really indistinct… owing to the ribosomes in the limiting membrane of the crystal, one could even assume that this crystal might be in the perinuclear space.





 .
.