Hepatocyte from a 14CoS -/- mouse (homozygous for a 3800kb deletion on chromosome)that has spent 360 days being rescued from certain demise by NTBC has little condensed chromatin, very round and active looking nucleus and literally thousands of nuclear pores, more than I remember seeing in the nuclear envelope of any other cell type, and any other experimental condition. Totally awesome. I would not hazard a guess as to how many are there per micron sq.
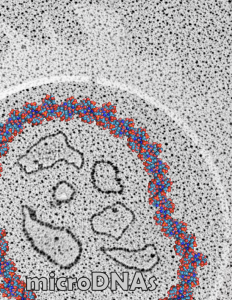
The purpose of nuclear pores is of course do provide a mechanism for selective transport of molecules in and out of the nucleus, separating the compartment of the cytoplasm and nucleus. All produced proteins from the cytoplasm required by the nucleus will be imported through the nuclear pore, and the RNAs, mRNA, tRNA and rRNA, destined to become cytoplasmic machinery, transcribed and modified (post-transcriptional modification) in the nucleus will be exported to the cytoplasm through nuclear pores.
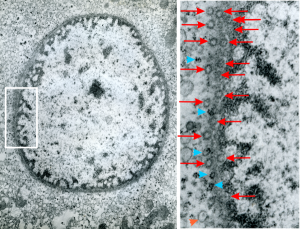
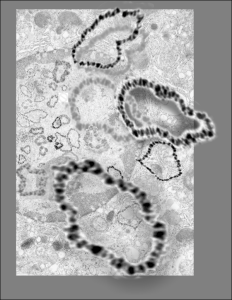
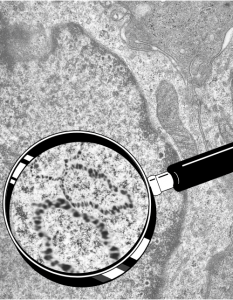
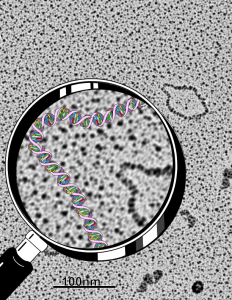
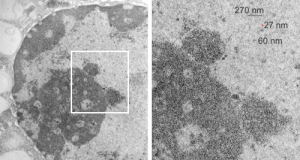
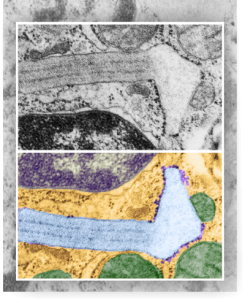
So the electron micrograph below (and inset) show this massive number of nuclear pores, particularly on one side that is cutting through the nuclear membrane at a slightly tangential angle, and also peculiarly, the polysomes (clear spirals of mRNA and rRNA) on the cytoplasmic side of the outer nuclear membrane are just about the only place in the cytoplasm where one finds RER. This vesicular ultrastructure (and particularly the vesicle – within – a vesicle) has been published Deiter et al 14cos, but not the nuclear pore observation. Here is such an hepatocyte nucleus, and inset to show tangentially cut nuclear pores (arrows), and spirals of mRNA with ribosomes (arrowheads). 17709_65053_14CoS-/- 360d w NTBC.