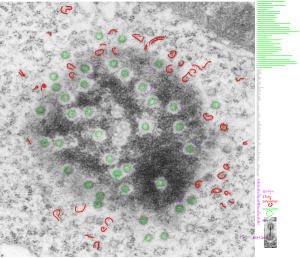
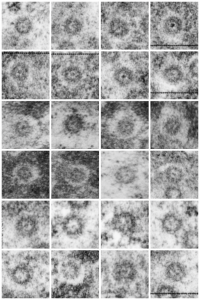
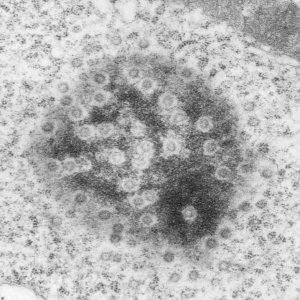
Adding more profiles of nuclear pores, from several different knock-out mice, basically to see whether the parameters I have set (chromatin exclusion zone, nuclear pore to nuclear pore distance, and distance between 20 nm DNA particles) changes with metabolic, or oxidative, or cell cycle (including apoptosis) stressors, and to see whether densities (probably pre-mRNA and other larger proteins that show up as occupants of the center of the nuclear pore) can be found to vary with similar cell states. I subjectively choose the nicer looking pores, tending to bet mid-cuts, and some subconscious bias likely causes me to choose those pores with transport molecules, as here about 87% of pores have center densities, while when counted in a single micrograph (not my selection) the ratio of pores with and without central densities is closer to 50%. we will see. The octagonal symmetry is very clear.