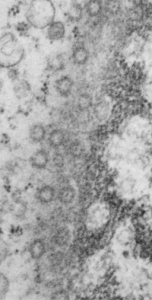
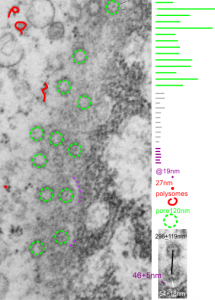
This hepatocyte was from a mouse (#5) which was a 14CoS nujll that did not receive NTBC as a rescue drug. This is 24 hours into the neonatal period. Nuclear pores have fewer central densities (presumably being proteins transported). I am measuring for distance beside the nuclear pore that the chromatin exclusion zone hoping that differences will show up with various conditions in previous experimental models.
Top micrograph, unretouched, bottom micrograph, pores used in calculation have circles, red dot = a 27nm ribosome, purple dots are areas of the chromatin just adjacent to the chromatin exclusion zone surrounding the nuclear pore.