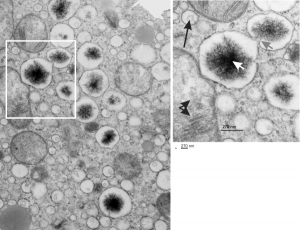
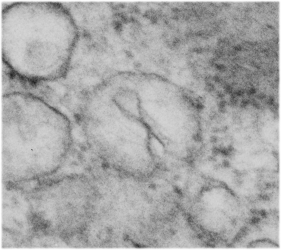
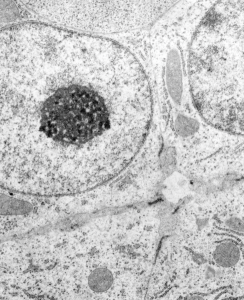
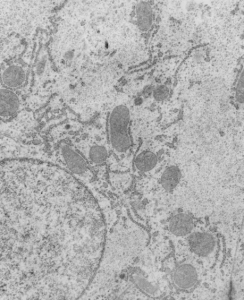
The four (maybe there are more) things that I noticed about the GCLC conditional KO mouse hepatocytes were 1: vesicles within vesicles, 2: lucent areas of mitochondiral matrix adjacent to quite well organized stacked cristi, 3: iron-like staining spicular deposits in dilated RER of hepatocytes, and 4: there is a linked-look between the ballooned areas of the ER (both smooth and rough).
These mice have survived the KO of GCLC, for 60 days, because they were rescued with N-acetyl-cysteine (NAC). NAC reportedly helps replenish the intracellular glutathione supply and can lessen hepatocyte oxidative damage occuring with the knock out of glutamate cysteine ligase catalytic subunit (Gclc), explained HERE. NAC affords protection against the oxidative stress, DNA damage and cancer development and cell death, thus rescues a genetically determined depletion of hepatocyte glutathione. I examined a large series of mouse tissues from these animals and their appropriate controls and while some of these data were published, there was much that never made it into the literature. Thus…. these posts, hoping that someone can be helped by these electron micrographic images of the unrescued, the controls and the NAC-saved mice.
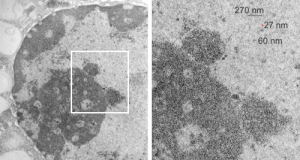
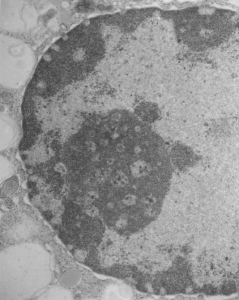
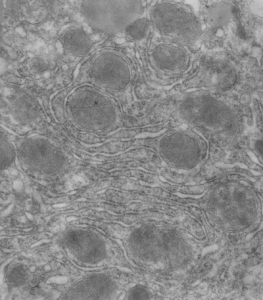
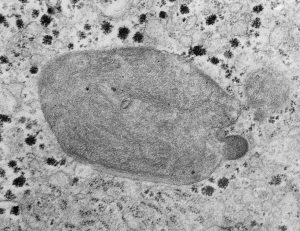
So the micrographs below have RER (albeit the ribosomes are really not that nicely arranged in spirals or grids) and the transition from RER to ER is not consistent with typical RER to SER transitions (that is; both small and large vesicles, empty and full of electron dense filaments; those with center vesicles and those without (almost like continuous segmented mini-intussuceptions, are kind of a large mix of different types, sizes, filled and empty vesicles. These vesicles begin to develope before D60 after NAC rescue, and increase with increasing time.
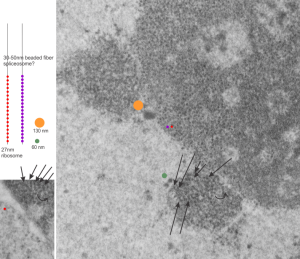
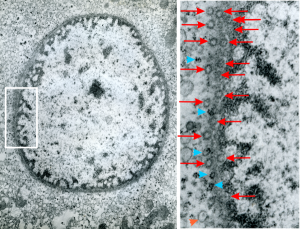
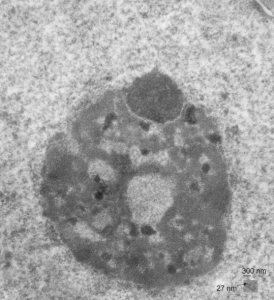
negative block and animal and treatment: 17902_74138_706_wcii_nac_60d. Size of a ribosome (at 27 nm) was used to calculate the bar markers (smallest black dot below. Inset from image on left is bounded with a white box, enlarge just around 2x on the right). Black arrow points to vesicle-within-a-vesicle phenomenon which is totally everywhere; double headed black arrow points to two types of mitochondrial inner membrane organization, one part points to areas which are quite electron-lucent, and one points to cristi that are highly layered and flattened. White arrow points to filamentous electron dense (i would like to say “iron filings” – like) inclusions and the grey arrow points to a telescoping point between two rounded contiguous vesicles with filamentous inclusions. Occasional ribosomes are seen on the bounding membranes of these vesicles, and as in this case, they look like peristaltic connections, dipping one end into the next rounded vesicle as if there were strictures, or boluses passing through a long undulating series of tubes. Thee are a few coiled ribosomes on an mRNA through out the cytoplasm, and not pictured some microbodies (dense bodies with uricase crystals). Image to right shows the open-closed RER, SER, ER profiles show internally.