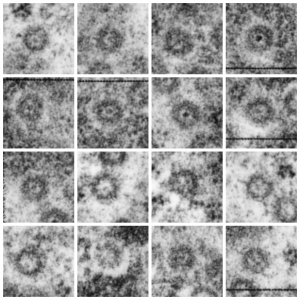
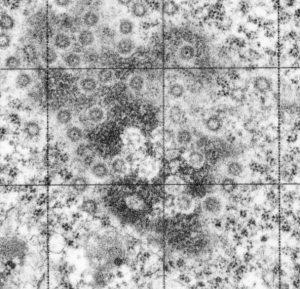
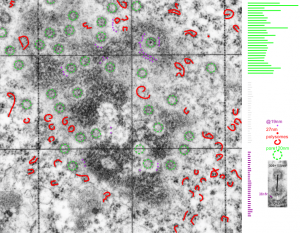
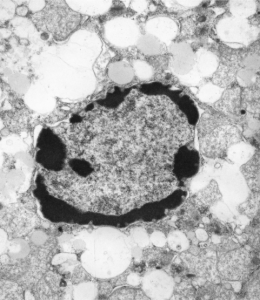
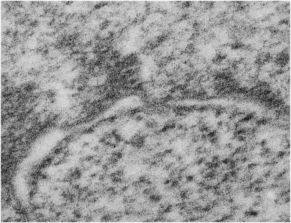
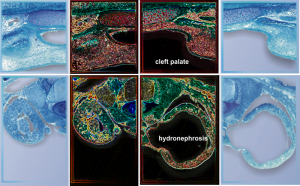
It sometimes overtakes me: Just the thought of the complexity of life as i see it through the electron microscope. An unbelievable maize of elements, most of which are still “unseen” and “unimagined” – one can spend a lifetime x 1000 building, searching, finding the patterns and puzzles of the ultrastructure of the cell.
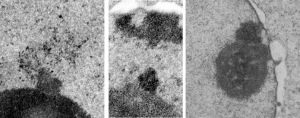
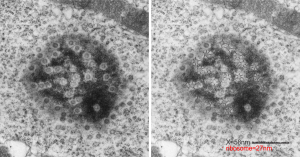
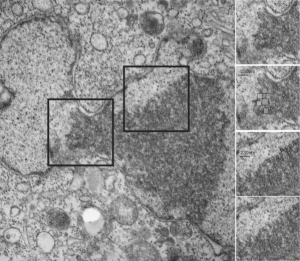
Considering the tangle of filaments and fibers and fibrils and proteins as “beads” like pre-RNA, RNA, or nucleosomes or wrapped up like balls of string, fractal tangles (tangles is not the right word of course-as they are only tangles if one can not undo-them) and recursive patterns both physical and in DNA code, sub-structures that layer in unusual ways (SP-A, langerin, or anti-parallel helix loop helix proteins just naming three) that can look like little arms reaching out from highly ordered nuclear pores where it is possible even to see pre-RNA densities moving through the central portion of the nuclear pore, and in which one can imagine smaller molecules floating through on the outer-but still-inner confines of those 120nm wide windows. Top down views of pores show rich octagonal symmetry (known by others to contain thousands of proteins), is extra special for the chromatin exclusion zone…. which provides little renaissance “arches” to protect, and to facilitate transport in and out of the nucleus.
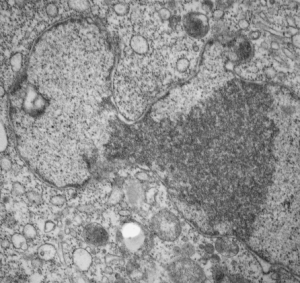
And with hilarity, a mitochondrion can have as many (probably more) as three little dips of its organellar body, with outer nuclear membrane visibly pulld down to proximate nuclear pores to become exclusively entangled with the outer pore filaments. And why is the nuclear basket on the inner nuclear membrane not that well defined in apoptosis? Is it that transport has become limited in cells preparing for self-destruct, closing up shop to preserve the well-being of the whole?
And this doesn’t speak yet to frequency of any of the very few selected items mentioned above as to mitochondria-cristi orientation and morphology, size and frequency; nuclear pores — frequency and morphology during high metabolic activity; and does not even begin to mention other organelles like golgi – size frequency, morphology, as perhaps, with the genetic absence of some magnesium transporter, or some sym or antiporters, channels, transport vesicles or influence of malfunction, or ER stress or environment… The whole thing is so complicated, so unknowable, so foreboding in its awesomeness and so frustrating to investigate, it just brings me to weep.